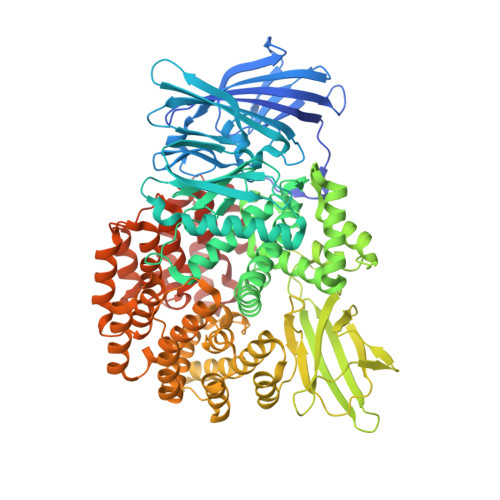
Structural basis for the unusual specificity of Escherichia coli aminopeptidase N.
Addlagatta, A., Gay, L., Matthews, B.W.(2008) Biochemistry 47: 5303-5311
- PubMed: 18416562 Search on PubMed
- DOI: https://doi.org/10.1021/bi7022333
- Primary Citation Related Structures:
3B2P, 3B2X, 3B34, 3B37, 3B3B - PubMed Abstract:
Aminopeptidase N from Escherichia coli is a M1 class aminopeptidase with the active-site region related to that of thermolysin. The enzyme has unusual specificity, cleaving adjacent to the large, nonpolar amino acids Phe and Tyr but also cleaving next to the polar residues Lys and Arg. To try to understand the structural basis for this pattern of hydrolysis, the structure of the enzyme was determined in complex with the amino acids L-arginine, L-lysine, L-phenylalanine, L-tryptophan, and L-tyrosine. These amino acids all bind with their backbone atoms close to the active-site zinc ion and their side chain occupying the S1 subsite. This subsite is in the form of a cylinder, about 10 A in cross-section and 12 A in length. The bottom of the cylinder includes the zinc ion and a number of polar side chains that make multiple hydrogen-bonding and other interactions with the alpha-amino group and the alpha-carboxylate of the bound amino acid. The walls of the S1 cylinder are hydrophobic and accommodate the nonpolar or largely nonpolar side chains of Phe and Tyr. The top of the cylinder is polar in character and includes bound water molecules. The epsilon-amino group of the bound lysine side chain and the guanidinium group of arginine both make multiple hydrogen bonds to this part of the S1 site. At the same time, the hydrocarbon part of the lysine and arginine side chains is accommodated within the nonpolar walls of the S1 cylinder. This combination of hydrophobic and hydrophilic binding surfaces explains the ability of ePepN to cleave Lys, Arg, Phe, and Tyr. Another favored substrate has Ala at the P1 position. The short, nonpolar side chain of this residue can clearly be bound within the hydrophobic part of the S1 cylinder, but the reason for its facile hydrolysis remains uncertain.
- Institute of Molecular Biology, Howard Hughes Medical Institute, and Department of Physics, 1229 University of Oregon, Eugene, Oregon 97403-1229, USA.
Organizational Affiliation: