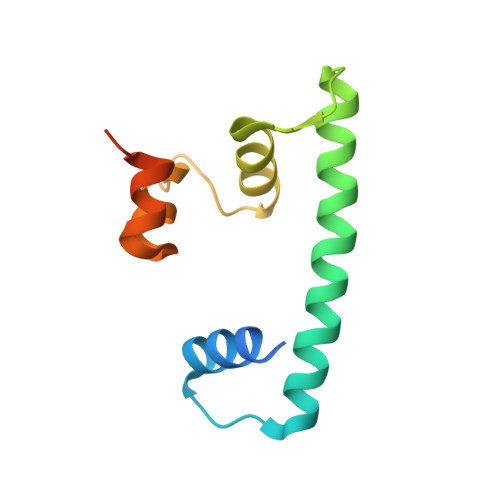
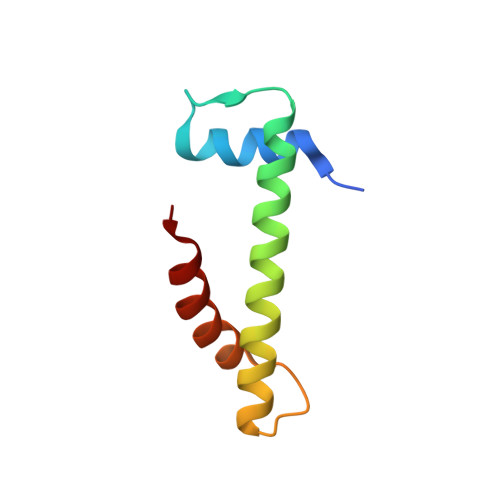
CENP-T-W-S-X Forms a Unique Centromeric Chromatin Structure with a Histone-like Fold.
Nishino, T., Takeuchi, K., Gascoigne, K.E., Suzuki, A., Hori, T., Oyama, T., Morikawa, K., Cheeseman, I.M., Fukagawa, T.(2012) Cell 148: 487-501
- PubMed: 22304917
- DOI: https://doi.org/10.1016/j.cell.2011.11.061
- Primary Citation Related Structures:
3B0B, 3B0C, 3B0D, 3VH5, 3VH6 - PubMed Abstract:
The multiprotein kinetochore complex must assemble at a specific site on each chromosome to achieve accurate chromosome segregation. Defining the nature of the DNA-protein interactions that specify the position of the kinetochore and provide a scaffold for kinetochore formation remain key goals. Here, we demonstrate that the centromeric histone-fold-containing CENP-T-W and CENP-S-X complexes coassemble to form a stable CENP-T-W-S-X heterotetramer. High-resolution structural analysis of the individual complexes and the heterotetramer reveals similarity to other histone fold-containing complexes including canonical histones within a nucleosome. The CENP-T-W-S-X heterotetramer binds to and supercoils DNA. Mutants designed to compromise heterotetramerization or the DNA-protein contacts around the heterotetramer strongly reduce the DNA binding and supercoiling activities in vitro and compromise kinetochore assembly in vivo. These data suggest that the CENP-T-W-S-X complex forms a unique nucleosome-like structure to generate contacts with DNA, extending the "histone code" beyond canonical nucleosome proteins.
- Department of Molecular Genetics, National Institute of Genetics and The Graduate University for Advanced Studies (SOKENDAI), Mishima, Shizuoka 411-8540, Japan.
Organizational Affiliation: