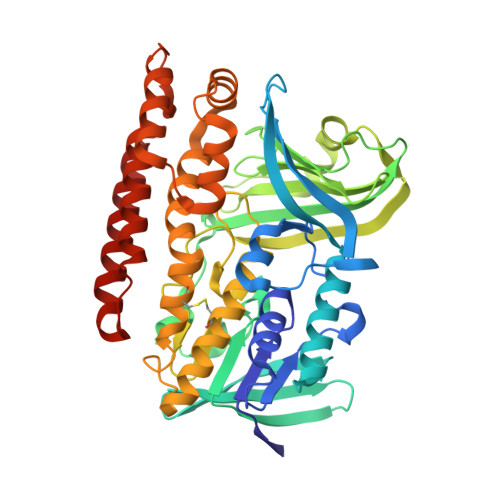
Structure and mutation analysis of archaeal geranylgeranyl reductase
Sasaki, D., Fujihashi, M., Iwata, Y., Murakami, M., Yoshimura, T., Hemmi, H., Miki, K.(2011) J Mol Biology 409: 543-557
- PubMed: 21515284 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.04.002
- Primary Citation Related Structures:
3ATQ, 3ATR - PubMed Abstract:
The crystal structure of geranylgeranyl reductase (GGR) from Sulfolobus acidocaldarius was determined in order to elucidate the molecular mechanism of the catalytic reaction. The enzyme is a flavoprotein and is involved in saturation of the double bonds on the isoprenoid moiety of archaeal membranes. The structure determined in this study belongs to the p-hydroxybenzoate hydroxylase family in the glutathione reductase superfamily. GGR functions as a monomer and is divided into the FAD-binding, catalytic and C-terminal domains. The catalytic domain has a large cavity surrounded by a characteristic YxWxFPx(7-8)GxG motif and by the isoalloxazine ring of an FAD molecule. The cavity holds a lipid molecule, which is probably derived from Escherichia coli cells used for over-expression. One of the two forms of the structure clarifies the presence of an anion pocket holding a pyrophosphate molecule, which might anchor the phosphate head of the natural ligands. Mutational analysis supports the suggestion that the three aromatic residues of the YxWxFPx(7-8)GxG motif hold the ligand in the appropriate position for reduction. Cys47, which is widely conserved in GGRs, is located at the si-side of the isoalloxazine ring of FAD and is shown by mutational analysis to be involved in catalysis. The catalytic cycle, including the FAD reducing factor binding site, is proposed on the basis of the detailed analysis of the structure.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto, Kyoto 606-8502, Japan.
Organizational Affiliation: