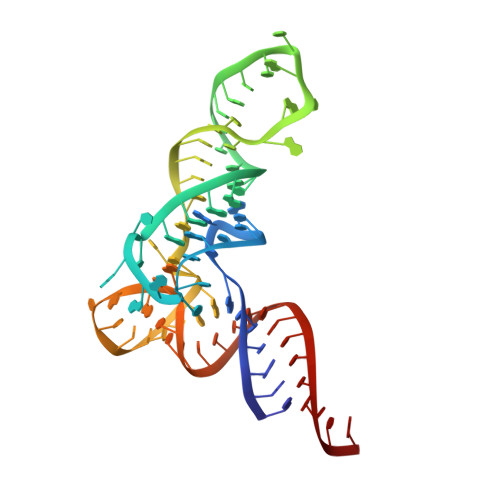
Structural basis of tRNA agmatinylation essential for AUA codon decoding
Osawa, T., Kimura, S., Terasaka, N., Inanaga, H., Suzuki, T., Numata, T.(2011) Nat Struct Mol Biol 18: 1275-1280
- PubMed: 22002223 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2144
- Primary Citation Related Structures:
3AMT, 3AMU, 3AU7 - PubMed Abstract:
The cytidine at the first position of the anticodon (C34) in the AUA codon-specific archaeal tRNA(Ile2) is modified to 2-agmatinylcytidine (agm(2)C or agmatidine), an agmatine-conjugated cytidine derivative, which is crucial for the precise decoding of the genetic code. Agm(2)C is synthesized by tRNA(Ile)-agm(2)C synthetase (TiaS) in an ATP-dependent manner. Here we present the crystal structures of the Archaeoglobus fulgidus TiaS-tRNA(Ile2) complexed with ATP, or with AMPCPP and agmatine, revealing a previously unknown kinase module required for activating C34 by phosphorylation, and showing the molecular mechanism by which TiaS discriminates between tRNA(Ile2) and tRNA(Met). In the TiaS-tRNA(Ile2)-ATP complex, C34 is trapped within a pocket far away from the ATP-binding site. In the agmatine-containing crystals, C34 is located near the AMPCPP γ-phosphate in the kinase module, demonstrating that agmatine is essential for placing C34 in the active site. These observations also provide the structural dynamics for agm(2)C formation.
- Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Ibaraki, Japan.
Organizational Affiliation: