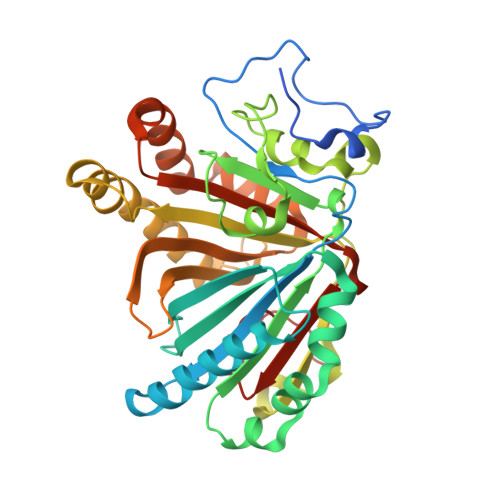
X-ray crystal structure of michaelis complex of aldoxime dehydratase
Sawai, H., Sugimoto, H., Kato, Y., Asano, Y., Shiro, Y., Aono, S.(2009) J Biological Chem 284: 32089-32096
- PubMed: 19740758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.018762
- Primary Citation Related Structures:
3A15, 3A16, 3A17, 3A18 - PubMed Abstract:
Aldoxime dehydratase (Oxd) catalyzes the dehydration of aldoximes (R-CH=N-OH) to their corresponding nitrile (R-C triple bond N). Oxd is a heme-containing enzyme that catalyzes the dehydration reaction as its physiological function. We have determined the first two structures of Oxd: the substrate-free OxdRE at 1.8 A resolution and the n-butyraldoxime- and propionaldoxime-bound OxdREs at 1.8 and 1.6 A resolutions, respectively. Unlike other heme enzymes, the organic substrate is directly bound to the heme iron in OxdRE. We determined the structure of the Michaelis complex of OxdRE by using the unique substrate binding and activity regulation properties of Oxd. The Michaelis complex was prepared by x-ray cryoradiolytic reduction of the ferric dead-end complex in which Oxd contains a Fe(3+) heme form. The crystal structures reveal the mechanism of substrate recognition and the catalysis of OxdRE.
- From the Okazaki Institute for Integrative Bioscience, National Institutes of Natural Sciences, 5-1 Higashiyama, Myodaiji, Okazaki 444-8787, Japan.
Organizational Affiliation: