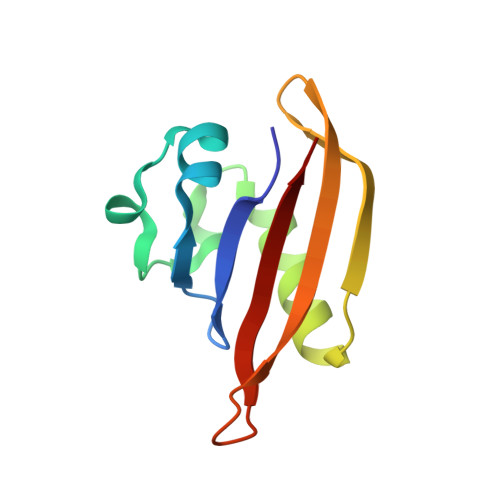
Structure of PAS-linked histidine kinase and the response regulator complex
Yamada, S., Sugimoto, H., Kobayashi, M., Ohno, A., Nakamura, H., Shiro, Y.(2009) Structure 17: 1333-1344
- PubMed: 19836334
- DOI: https://doi.org/10.1016/j.str.2009.07.016
- Primary Citation of Related Structures:
3A0R, 3A0S, 3A0T, 3A0U, 3A0V, 3A0W, 3A0X, 3A0Y, 3A0Z, 3A10 - PubMed Abstract:
We determined the structure of the complex of the sensory histidine kinase (HK) and its cognate response regulator (RR) in the two-component signal transduction system of Thermotoga maritima. This was accomplished by fitting the high-resolution structures of the isolated HK domains and the RR onto the electron density map (3.8 A resolution) of the HK/RR complex crystal. Based on the structural information, we evaluated the roles of both interdomain and intermolecular interactions in the signal transduction of the cytosolic PAS-linked HK and RR system, in particular the O(2)-sensor FixL/FixJ system. The PAS-sensor domain of HK interacts with the catalytic domain of the same polypeptide chain by creating an interdomain beta sheet. The interaction site between HK and RR, which was confirmed by NMR, is suitable for the intermolecular transfer reaction of the phosphoryl group, indicating that the observed interaction is important for the phosphatase activity of HK that dephosphorylates phospho-RR.
- RIKEN SPring-8 Center, Kouto 1-1-1, Sayo, Hyogo, Japan.
Organizational Affiliation: