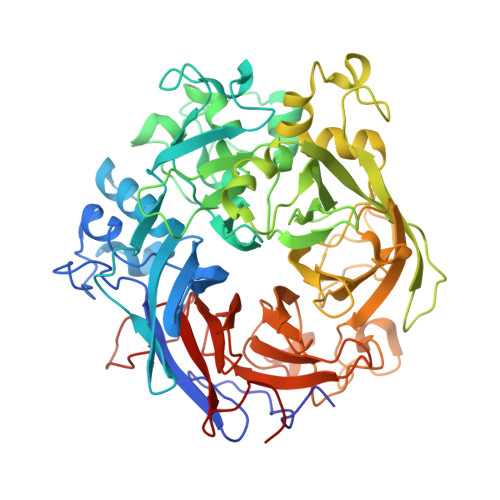
A Complex Iron-Calcium Cofactor Catalyzing Phosphotransfer Chemistry
Yong, S.C., Roversi, P., Lillington, J., Rodriguez, F., Krehenbrink, M., Zeldin, O.B., Garman, E.F., Lea, S.M., Berks, B.C.(2014) Science 345: 1170
- PubMed: 25190793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1254237
- Primary Citation Related Structures:
3ZWU, 4A9V, 4A9X, 4ALF, 4AMF - PubMed Abstract:
Alkaline phosphatases play a crucial role in phosphate acquisition by microorganisms. To expand our understanding of catalysis by this class of enzymes, we have determined the structure of the widely occurring microbial alkaline phosphatase PhoX. The enzyme contains a complex active-site cofactor comprising two antiferromagnetically coupled ferric iron ions (Fe(3+)), three calcium ions (Ca(2+)), and an oxo group bridging three of the metal ions. Notably, the main part of the cofactor resembles synthetic oxide-centered triangular metal complexes. Structures of PhoX-ligand complexes reveal how the active-site metal ions bind substrate and implicate the cofactor oxo group in the catalytic mechanism. The presence of iron in PhoX raises the possibility that iron bioavailability limits microbial phosphate acquisition.
- Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: