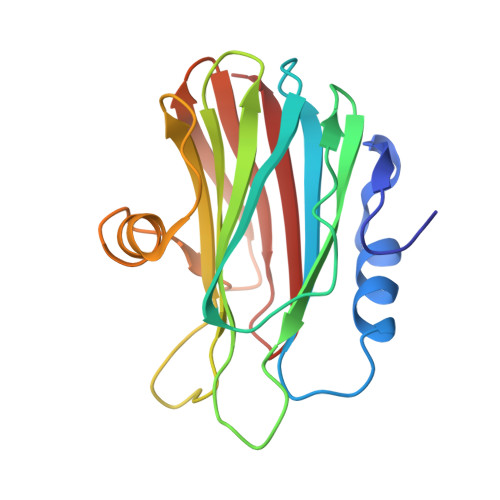
Pores of the Toxin Frac Assemble Into 2D Hexagonal Clusters in Both Crystal Structures and Model Membranes.
Mechaly, A.E., Bellomio, A., Morante, K., Agirre, J., Gil-Carton, D., Valle, M., Gonzalez-Manas, J.M., Guerin, D.M.A.(2012) J Struct Biol 180: 312
- PubMed: 22728830 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2012.06.003
- Primary Citation Related Structures:
3ZWG, 3ZWJ - PubMed Abstract:
The recent high-resolution structure of the toxin FraC derived from the sea anemone Actinia fragacea has provided new insight into the mechanism of pore formation by actinoporins. In this work, we report two new crystal forms of FraC in its oligomeric prepore conformation. Together with the previously reported structure, these two new structures reveal that ring-like nonamers of the toxin assemble into compact two-dimensional hexagonal arrays. This supramolecular organization is maintained in different relative orientations adopted by the oligomers within the crystal layers. Analyses of the aggregation of FraC pores in both planar and curved (vesicles) model membranes show similar 2D hexagonal arrangements. Our observations support a model in which hexagonal pore-packing is a clustering mechanism that maximizes toxin-driven membrane damage in the target cell.
- Unidad de Biofísica (Centro Mixto CSIC-UPV/EHU), B Sarriena S/N, Campus UPV/EHU Leioa, Leioa, Vizcaya, Spain.
Organizational Affiliation: