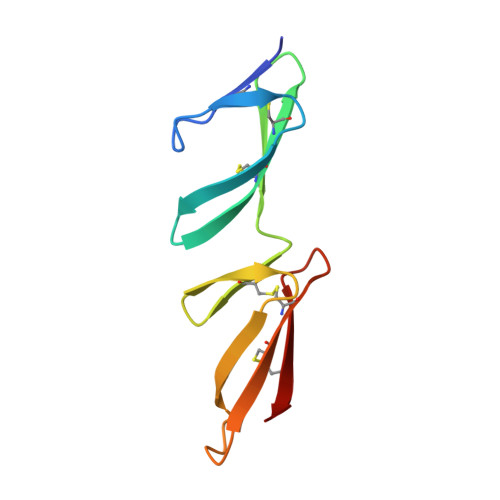
Structural and Functional Analysis of the Tandem Beta-Zipper Interaction of a Streptococcal Protein with Human Fibronectin.
Norris, N.C., Bingham, R.J., Harris, G., Speakman, A., Jones, R.P.O., Leech, A., Turkenburg, J.P., Potts, J.R.(2011) J Biological Chem 286: 38311
- PubMed: 21840989 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.276592
- Primary Citation Related Structures:
3ZRZ - PubMed Abstract:
Bacterial fibronectin-binding proteins (FnBPs) contain a large intrinsically disordered region (IDR) that mediates adhesion of bacteria to host tissues, and invasion of host cells, through binding to fibronectin (Fn). These FnBP IDRs consist of Fn-binding repeats (FnBRs) that form a highly extended tandem β-zipper interaction on binding to the N-terminal domain of Fn. Several FnBR residues are highly conserved across bacterial species, and here we investigate their contribution to the interaction. Mutation of these residues to alanine in SfbI-5 (a disordered FnBR from the human pathogen Streptococcus pyogenes) reduced binding, but for each residue the change in free energy of binding was <2 kcal/mol. The structure of an SfbI-5 peptide in complex with the second and third F1 modules from Fn confirms that the conserved FnBR residues play equivalent functional roles across bacterial species. Thus, in SfbI-5, the binding energy for the tandem β-zipper interaction with Fn is distributed across the interface rather than concentrated in a small number of "hot spot" residues that are frequently observed in the interactions of folded proteins. We propose that this might be a common feature of the interactions of IDRs and is likely to pose a challenge for the development of small molecule inhibitors of FnBP-mediated adhesion to and invasion of host cells.
- Department of Biology, University of York, York, YO10 5DD, United Kingdom.
Organizational Affiliation: