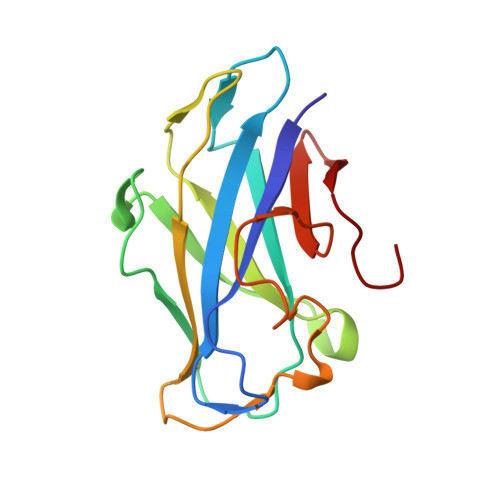
A Single Mutation Reforms the Binding Activity of an Adhesion-Deficient Family 3 Carbohydrate-Binding Module
Yaniv, O., Petkun, S., Shimon, L.J.W., Bayer, E.A., Lamed, R., Frolow, F.(2012) Acta Crystallogr D Biol Crystallogr 68: 819
- PubMed: 22751667 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912013133
- Primary Citation Related Structures:
2YLK, 3ZQX - PubMed Abstract:
The crystal structure of the family 3b carbohydrate-binding module (CBM3b) of the cellulosomal multimodular hydrolytic enzyme cellobiohydrolase 9A (Cbh9A) from Clostridium thermocellum has been determined. Cbh9A CBM3b crystallized in space group P4(1) with four molecules in the asymmetric unit and diffracted to a resolution of 2.20 Å using synchrotron radiation. The structure was determined by molecular replacement using C. thermocellum Cel9V CBM3b' (PDB entry 2wnx) as a model. The C. thermocellum Cbh9A CBM3b molecule forms a nine-stranded antiparallel β-sandwich similar to other family 3 carbohydrate-binding modules (CBMs). It has a short planar array of two aromatic residues that are assumed to bind cellulose, yet it lacks the ability to bind cellulose. The molecule contains a shallow groove of unknown function that characterizes other family 3 CBMs with high sequence homology. In addition, it contains a calcium-binding site formed by a group of amino-acid residues that are highly conserved in similar structures. After determination of the three-dimensional structure of Cbh9A CBM3b, the site-specific N126W mutant was produced with the intention of enhancing the cellulose-binding ability of the CBM. Cbh9A CBM3b(N126W) crystallized in space group P4(1)2(1)2, with one molecule in the asymmetric unit. The crystals diffracted to 1.04 Å resolution using synchrotron radiation. The structure of Cbh9A CBM3b(N126W) revealed incorporation of the mutated Trp126 into the putative cellulose-binding strip of residues. Cellulose-binding experiments demonstrated the ability of Cbh9A CBM3b(N126W) to bind cellulose owing to the mutation. This is the first report of the engineered conversion of a non-cellulose-binding CBM3 to a binding CBM3 by site-directed mutagenesis. The three-dimensional structure of Cbh9A CBM3b(N126W) provided a structural correlation with cellulose-binding ability, revealing a longer planar array of definitive cellulose-binding residues.
- Department of Molecular Microbiology and Biotechnology, Tel Aviv University, Tel Aviv 69978, Israel.
Organizational Affiliation: