On Terminal Alkynes that Can React with Active-Site Cysteine Nucleophiles in Proteases.
Ekkebus, R., Van Kasteren, S.I., Kulathu, Y., Scholten, A., Berlin, I., Geurink, P.P., De Jong, A., Goerdayal, G., Neefjes, J., Heck, A.J.R., Komander, D., Ovaa, H.(2013) J Am Chem Soc 135: 2867
- PubMed: 23387960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja309802n
- Primary Citation Related Structures:
3ZNH - PubMed Abstract:
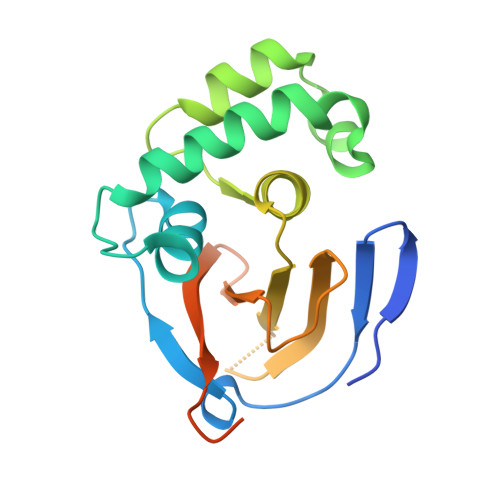
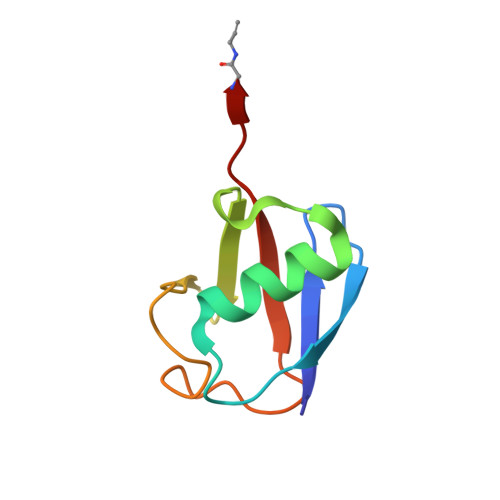
Active-site directed probes are powerful in studies of enzymatic function. We report an active-site directed probe based on a warhead so far considered unreactive. By replacing the C-terminal carboxylate of ubiquitin (Ub) with an alkyne functionality, a selective reaction with the active-site cysteine residue of de-ubiquitinating enzymes was observed. The resulting product was shown to be a quaternary vinyl thioether, as determined by X-ray crystallography. Proteomic analysis of proteins bound to an immobilized Ub alkyne probe confirmed the selectivity toward de-ubiquitinating enzymes. The observed reactivity is not just restricted to propargylated Ub, as highlighted by the selective reaction between caspase-1 (interleukin converting enzyme) and a propargylated peptide derived from IL-1β, a caspase-1 substrate.
- Division of Cell Biology, The Netherlands Cancer Institute, Amsterdam, The Netherlands.
Organizational Affiliation: