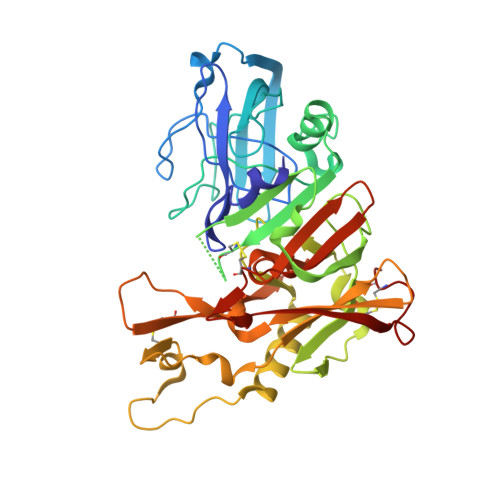
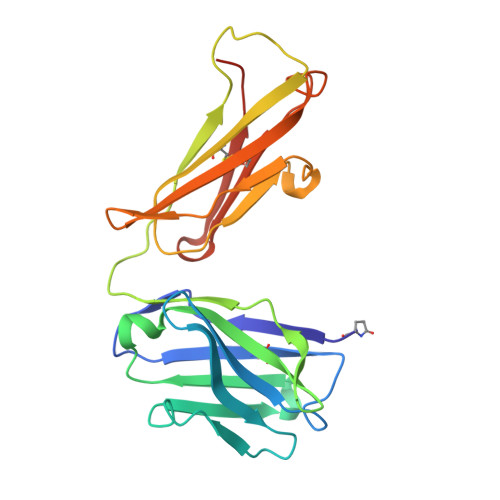
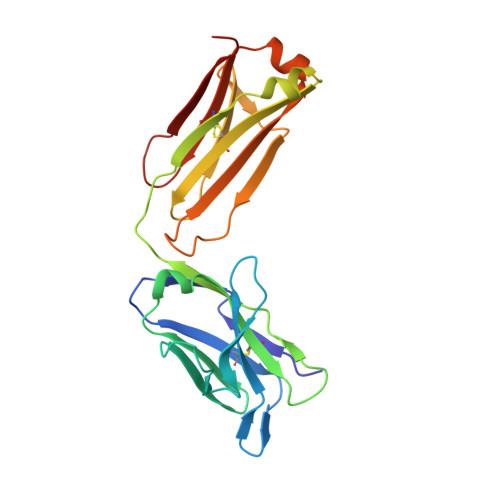
Mapping the Conformational Space Accessible to Bace2 Using Surface Mutants and Co-Crystals with Fab-Fragments, Fynomers, and Xaperones
Banner, D.W., Gsell, B., Benz, J., Bertschinger, J., Burger, D., Brack, S., Cuppuleri, S., Debulpaep, M., Gast, A., Grabulovski, D., Hennig, M., Hilpert, H., Huber, W., Kuglstatter, A., Kusznir, E., Laeremans, T., Matile, H., Miscenic, C., Rufer, A., Schlatter, D., Steyeart, J., Stihle, M., Thoma, R., Weber, M., Ruf, A.(2013) Acta Crystallogr D Biol Crystallogr 69: 1124
- PubMed: 23695257 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913006574
- Primary Citation Related Structures:
3ZKG, 3ZKI, 3ZKM, 3ZKN, 3ZKQ, 3ZKS, 3ZKX, 3ZL7, 3ZOV, 4BEL, 4BFB - PubMed Abstract:
The aspartic protease BACE2 is responsible for the shedding of the transmembrane protein Tmem27 from the surface of pancreatic β-cells, which leads to inactivation of the β-cell proliferating activity of Tmem27. This role of BACE2 in the control of β-cell maintenance suggests BACE2 as a drug target for diabetes. Inhibition of BACE2 has recently been shown to lead to improved control of glucose homeostasis and to increased insulin levels in insulin-resistant mice. BACE2 has 52% sequence identity to the well studied Alzheimer's disease target enzyme β-secretase (BACE1). High-resolution BACE2 structures would contribute significantly to the investigation of this enzyme as either a drug target or anti-target. Surface mutagenesis, BACE2-binding antibody Fab fragments, single-domain camelid antibody VHH fragments (Xaperones) and Fyn-kinase-derived SH3 domains (Fynomers) were used as crystallization helpers to obtain the first high-resolution structures of BACE2. Eight crystal structures in six different packing environments define an ensemble of low-energy conformations available to the enzyme. Here, the different strategies used for raising and selecting BACE2 binders for cocrystallization are described and the crystallization success, crystal quality and the time and resources needed to obtain suitable crystals are compared.
- pRED Pharma Research and Early Development, Small Molecule Research, Discovery Technologies, F. Hoffmann-La Roche Ltd, 4070 Basel, Switzerland.
Organizational Affiliation: