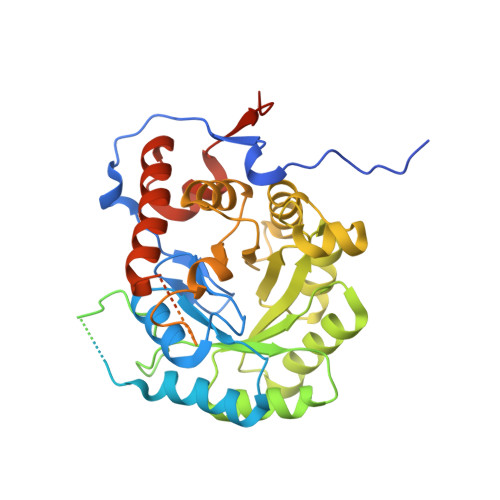
Structure of Pseudomonas Aeruginosa Inosine 5'-Monophosphate Dehydrogenase
Rao, V.A., Shepherd, S.M., Owen, R., Hunter, W.N.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 243
- PubMed: 23519796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113002352
- Primary Citation Related Structures:
3ZFH - PubMed Abstract:
Inosine 5'-monophosphate dehydrogenase (IMPDH) represents a potential antimicrobial drug target. The crystal structure of recombinant Pseudomonas aeruginosa IMPDH has been determined to a resolution of 2.25 Å. The structure is a homotetramer of subunits dominated by a (β/α)8-barrel fold, consistent with other known structures of IMPDH. Also in common with previous work, the cystathionine β-synthase domains, residues 92-204, are not present in the model owing to disorder. However, unlike the majority of available structures, clearly defined electron density exists for a loop that creates part of the active site. This loop, composed of residues 297-315, links α8 and β9 and carries the catalytic Cys304. P. aeruginosa IMPDH shares a high level of sequence identity with bacterial and protozoan homologues, with residues involved in binding substrate and the NAD+ cofactor being conserved. Specific differences that have been proven to contribute to selectivity against the human enzyme in a study of Cryptosporidium parvum IMPDH are also conserved, highlighting the potential value of IMPDH as a drug target.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dow Street, Dundee DD1 5EH, Scotland.
Organizational Affiliation: