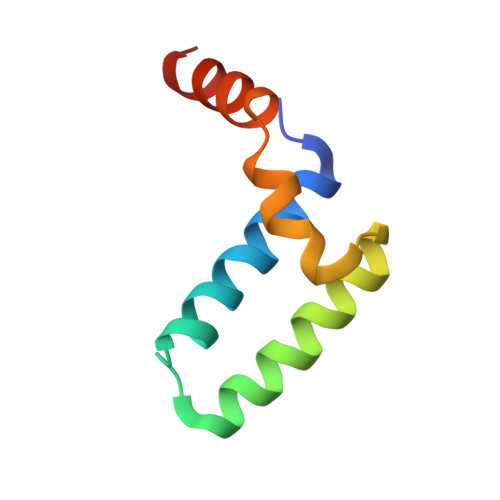
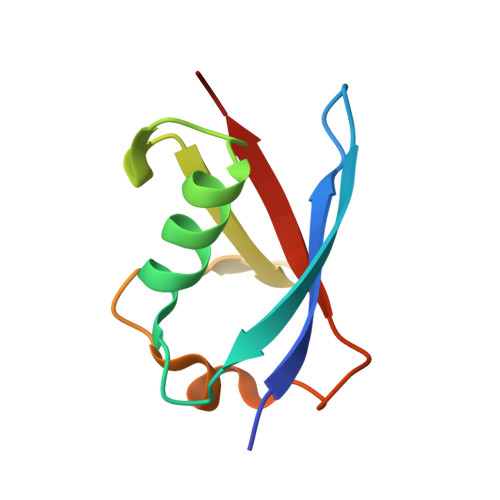
Structure of the Sgt2 Dimerization Domain Complexed with the Get5 Ubl Domain Involved in the Targeting of Tail-Anchored Membrane Proteins to the Endoplasmic Reticulum.
Tung, J.-Y., Li, Y.-C., Lin, T.-W., Hsiao, C.-D.(2013) Acta Crystallogr D Biol Crystallogr 69: 2081
- PubMed: 24100326 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913019379
- Primary Citation Related Structures:
3ZDM - PubMed Abstract:
The insertion of tail-anchored membrane (TA) proteins into the appropriate membrane is a post-translational event that requires stabilization of the transmembrane domain and targeting to the proper destination. Sgt2, a small glutamine-rich tetratricopeptide-repeat protein, is a heat-shock protein cognate (HSC) co-chaperone that preferentially binds endoplasmic reticulum-destined TA proteins and directs them to the GET pathway via Get4 and Get5. The N-terminal domain of Sgt2 seems to exert dual functions. It mediates Get5 interaction and allows substrate delivery to Get3. Following the N-terminus of Get5 is a ubiquitin-like (Ubl) domain that interacts with the N-terminus of Sgt2. Here, the crystal structure of the Sgt2 dimerization domain complexed with the Get5 Ubl domain (Sgt2N-Get5Ubl) is reported. This complex reveals an intimate interaction between one Sgt2 dimer and one Get5 monomer. This research further demonstrates that hydrophobic residues from both Sgt2 and Get5 play an important role in cell survival under heat stress. This study provides detailed molecular insights into the specific binding of this GET-pathway complex.
- Institute of Molecular Biology, Academia Sinica, Taipei 115, Taiwan.
Organizational Affiliation: