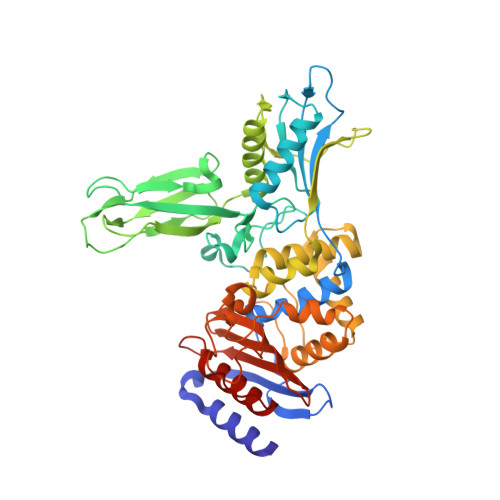
Inhibition of Dd-Peptidases by a Specific Trifluoroketone: Crystal Structure of a Complex with the Actinomadura R39 Dd-Peptidase.
Dzhekieva, L., Adediran, S.A., Herman, R., Kerff, F., Duez, C., Charlier, P., Sauvage, E., Pratt, R.F.(2013) Biochemistry 52: 2128
- PubMed: 23484909 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi400048s
- Primary Citation Related Structures:
3ZCZ - PubMed Abstract:
Inhibitors of bacterial DD-peptidases represent potential antibiotics. In the search for alternatives to β-lactams, we have investigated a series of compounds designed to generate transition state analogue structures upon reaction with DD-peptidases. The compounds contain a combination of a peptidoglycan-mimetic specificity handle and a warhead capable of delivering a tetrahedral anion to the enzyme active site. The latter includes a boronic acid, two alcohols, an aldehyde, and a trifluoroketone. The compounds were tested against two low-molecular mass class C DD-peptidases. As expected from previous observations, the boronic acid was a potent inhibitor, but rather unexpectedly from precedent, the trifluoroketone [D-α-aminopimelyl(1,1,1-trifluoro-3-amino)butan-2-one] was also very effective. Taking into account competing hydration, we found the trifluoroketone was the strongest inhibitor of the Actinomadura R39 DD-peptidase, with a subnanomolar (free ketone) inhibition constant. A crystal structure of the complex between the trifluoroketone and the R39 enzyme showed that a tetrahedral adduct had indeed formed with the active site serine nucleophile. The trifluoroketone moiety, therefore, should be considered along with boronic acids and phosphonates as a warhead that can be incorporated into new and effective DD-peptidase inhibitors and therefore, perhaps, antibiotics.
- Department of Chemistry, Wesleyan University , Lawn Avenue, Middletown, Connecticut 06459, United States.
Organizational Affiliation: