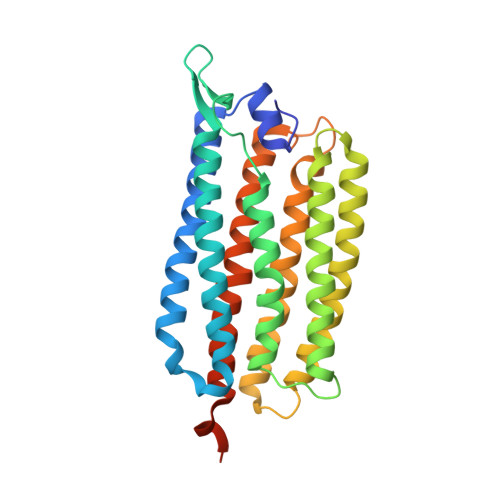
Structural basis for Na(+) transport mechanism by a light-driven Na(+) pump
Kato, H.E., Inoue, K., Abe-Yoshizumi, R., Kato, Y., Ono, H., Konno, M., Hososhima, S., Ishizuka, T., Hoque, M.R., Kunitomo, H., Ito, J., Yoshizawa, S., Yamashita, K., Takemoto, M., Nishizawa, T., Taniguchi, R., Kogure, K., Maturana, A.D., Iino, Y., Yawo, H., Ishitani, R., Kandori, H., Nureki, O.(2015) Nature 521: 48-53
- PubMed: 25849775 Search on PubMed
- DOI: https://doi.org/10.1038/nature14322
- Primary Citation Related Structures:
3X3B, 3X3C - PubMed Abstract:
Krokinobacter eikastus rhodopsin 2 (KR2) is the first light-driven Na(+) pump discovered, and is viewed as a potential next-generation optogenetics tool. Since the positively charged Schiff base proton, located within the ion-conducting pathway of all light-driven ion pumps, was thought to prohibit the transport of a non-proton cation, the discovery of KR2 raised the question of how it achieves Na(+) transport. Here we present crystal structures of KR2 under neutral and acidic conditions, which represent the resting and M-like intermediate states, respectively. Structural and spectroscopic analyses revealed the gating mechanism, whereby the flipping of Asp116 sequesters the Schiff base proton from the conducting pathway to facilitate Na(+) transport. Together with the structure-based engineering of the first light-driven K(+) pumps, electrophysiological assays in mammalian neurons and behavioural assays in a nematode, our studies reveal the molecular basis for light-driven non-proton cation pumps and thus provide a framework that may advance the development of next-generation optogenetics.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: