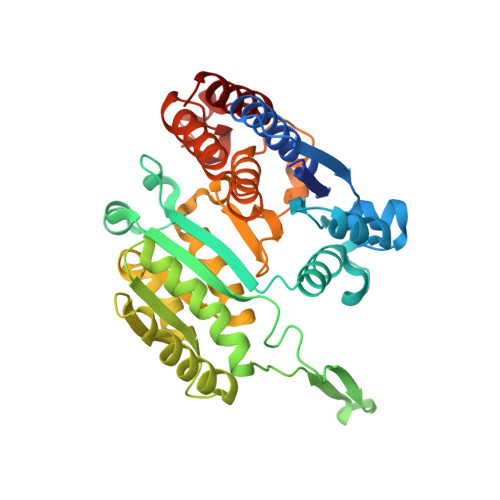
Pressure Adaptation of 3-Isopropylmalate Dehydrogenase from the Extremely Piezophilic Bacterium Shewanella benthica is Attributed to just One Amino Acid Substitution
Hamajima, Y., Nagae, T., Watanabe, N., Kato-Yamada, Y., Imai, T., Kato, C.To be published.