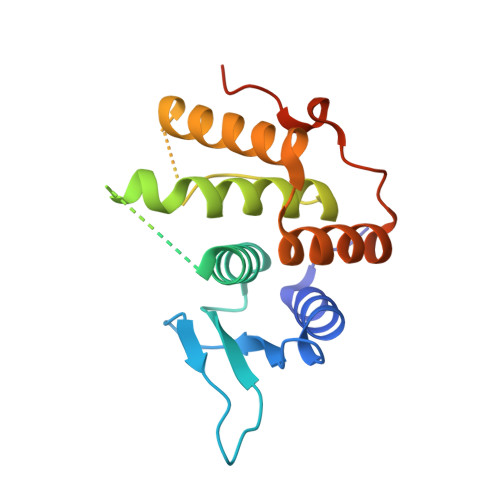
The non-canonical hydroxylase structure of YfcM reveals a metal ion-coordination motif required for EF-P hydroxylation
Kobayashi, K., Katz, A., Rajkovic, A., Ishii, R., Branson, O.E., Freitas, M.A., Ishitani, R., Ibba, M., Nureki, O.(2014) Nucleic Acids Res 42: 12295-12305
- PubMed: 25274739 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku898
- Primary Citation Related Structures:
3WTR, 4PDN - PubMed Abstract:
EF-P is a bacterial tRNA-mimic protein, which accelerates the ribosome-catalyzed polymerization of poly-prolines. In Escherichia coli, EF-P is post-translationally modified on a conserved lysine residue. The post-translational modification is performed in a two-step reaction involving the addition of a β-lysine moiety and the subsequent hydroxylation, catalyzed by PoxA and YfcM, respectively. The β-lysine moiety was previously shown to enhance the rate of poly-proline synthesis, but the role of the hydroxylation is poorly understood. We solved the crystal structure of YfcM and performed functional analyses to determine the hydroxylation mechanism. In addition, YfcM appears to be structurally distinct from any other hydroxylase structures reported so far. The structure of YfcM is similar to that of the ribonuclease YbeY, even though they do not share sequence homology. Furthermore, YfcM has a metal ion-coordinating motif, similar to YbeY. The metal ion-coordinating motif of YfcM resembles a 2-His-1-carboxylate motif, which coordinates an Fe(II) ion and forms the catalytic site of non-heme iron enzymes. Our findings showed that the metal ion-coordinating motif of YfcM plays an essential role in the hydroxylation of the β-lysylated lysine residue of EF-P. Taken together, our results suggested the potential catalytic mechanism of hydroxylation by YfcM.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan Global Research Cluster, RIKEN, 2-1, Hirosawa, Wako, Saitama, 351-0198, Japan.
Organizational Affiliation: