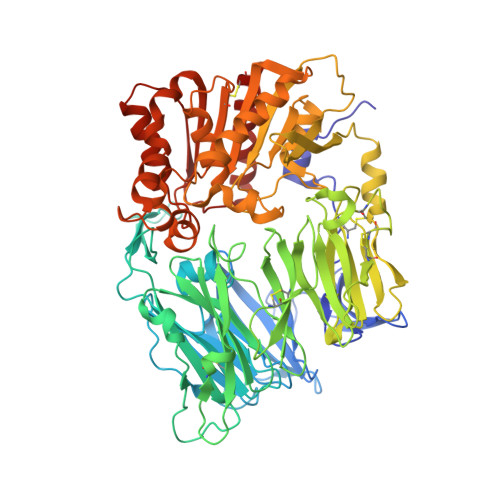
Anagliptin, a potent dipeptidyl peptidase IV inhibitor: its single-crystal structure and enzyme interactions.
Watanabe, Y.S., Yasuda, Y., Kojima, Y., Okada, S., Motoyama, T., Takahashi, R., Oka, M.(2015) J Enzyme Inhib Med Chem 30: 981-988
- PubMed: 26147347 Search on PubMed
- DOI: https://doi.org/10.3109/14756366.2014.1002402
- Primary Citation Related Structures:
3WQH - PubMed Abstract:
The single-crystal structure of anagliptin, N-[2-({2-[(2S)-2-cyanopyrrolidin-1-yl]-2-oxoethyl}amino)-2-methylpropyl]-2-methylpyrazolo[1,5-a]pyrimidine-6-carboxamide, was determined. Two independent molecules were held together by intermolecular hydrogen bonds, and the absolute configuration of the 2-cyanopyrrolidine ring delivered from l-prolinamide was confirmed to be S. The interactions of anagliptin with DPP-4 were clarified by the co-crystal structure solved at 2.85 Å resolution. Based on the structure determined by X-ray crystallography, the potency and selectivity of anagliptin were discussed, and an SAR study using anagliptin derivatives was performed.
- a Mie Research Park, Sanwa Kagaku Kenkyusho Co., Ltd. , Mie , Japan .
Organizational Affiliation: