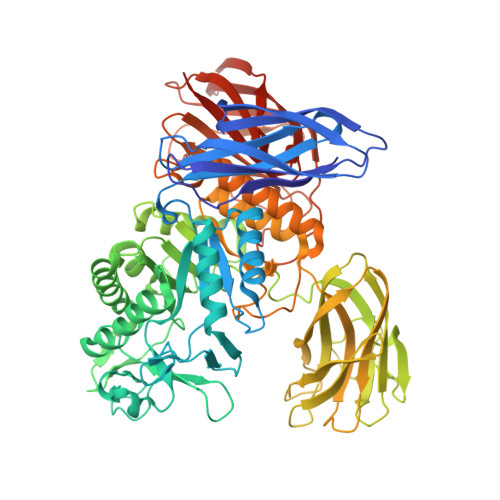
Structural elucidation of the cyclization mechanism of alpha-1,6-glucan by Bacillus circulans T-3040 cycloisomaltooligosaccharide glucanotransferase.
Suzuki, N., Fujimoto, Z., Kim, Y.M., Momma, M., Kishine, N., Suzuki, R., Suzuki, S., Kitamura, S., Kobayashi, M., Kimura, A., Funane, K.(2014) J Biological Chem 289: 12040-12051
- PubMed: 24616103 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.547992
- Primary Citation Related Structures:
3WNK, 3WNL, 3WNM, 3WNN, 3WNO - PubMed Abstract:
Bacillus circulans T-3040 cycloisomaltooligosaccharide glucanotransferase belongs to the glycoside hydrolase family 66 and catalyzes an intramolecular transglucosylation reaction that produces cycloisomaltooligosaccharides from dextran. The crystal structure of the core fragment from Ser-39 to Met-738 of B. circulans T-3040 cycloisomaltooligosaccharide glucanotransferase, devoid of its N-terminal signal peptide and C-terminal nonconserved regions, was determined. The structural model contained one catalytic (β/α)8-barrel domain and three β-domains. Domain N with an immunoglobulin-like β-sandwich fold was attached to the N terminus; domain C with a Greek key β-sandwich fold was located at the C terminus, and a carbohydrate-binding module family 35 (CBM35) β-jellyroll domain B was inserted between the 7th β-strand and the 7th α-helix of the catalytic domain A. The structures of the inactive catalytic nucleophile mutant enzyme complexed with isomaltohexaose, isomaltoheptaose, isomaltooctaose, and cycloisomaltooctaose revealed that the ligands bound in the catalytic cleft and the sugar-binding site of CBM35. Of these, isomaltooctaose bound in the catalytic site extended to the second sugar-binding site of CBM35, which acted as subsite -8, representing the enzyme·substrate complex when the enzyme produces cycloisomaltooctaose. The isomaltoheptaose and cycloisomaltooctaose bound in the catalytic cleft with a circular structure around Met-310, representing the enzyme·product complex. These structures collectively indicated that CBM35 functions in determining the size of the product, causing the predominant production of cycloisomaltooctaose by the enzyme. The canonical sugar-binding site of CBM35 bound the mid-part of isomaltooligosaccharides, indicating that the original function involved substrate binding required for efficient catalysis.
- Biomolecular Research Unit, National Institute of Agrobiological Sciences, Tsukuba 305-8602.
Organizational Affiliation: