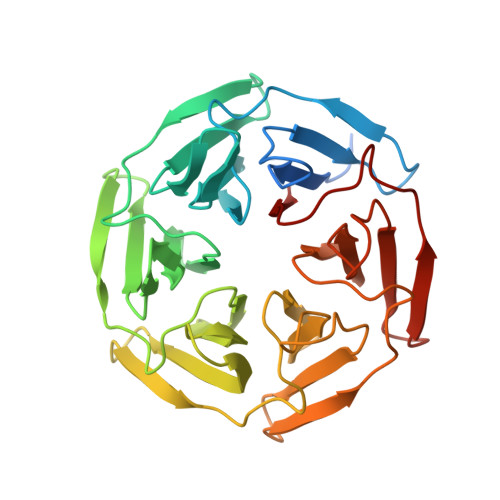
Kinetic, thermodynamic, and structural characterizations of the association between Nrf2-DLGex degron and Keap1
Fukutomi, T., Takagi, K., Mizushima, T., Ohuchi, N., Yamamoto, M.(2014) Mol Cell Biol 34: 832-846
- PubMed: 24366543
- DOI: https://doi.org/10.1128/MCB.01191-13
- Primary Citation Related Structures:
3WN7 - PubMed Abstract:
Transcription factor Nrf2 (NF-E2-related factor 2) coordinately regulates cytoprotective gene expression, but under unstressed conditions, Nrf2 is degraded rapidly through Keap1 (Kelch-like ECH-associated protein 1)-mediated ubiquitination. Nrf2 harbors two Keap1-binding motifs, DLG and ETGE. Interactions between these two motifs and Keap1 constitute a key regulatory nexus for cellular Nrf2 activity through the formation of a two-site binding hinge-and-latch mechanism. In this study, we determined the minimum Keap1-binding sequence of the DLG motif, the low-affinity latch site, and defined a new DLGex motif that covers a sequence much longer than that previously defined. We have successfully clarified the crystal structure of the Keap1-DC-DLGex complex at 1.6 Å. DLGex possesses a complicated helix structure, which interprets well the human-cancer-derived loss-of-function mutations in DLGex. In thermodynamic analyses, Keap1-DLGex binding is characterized as enthalpy and entropy driven, while Keap1-ETGE binding is characterized as purely enthalpy driven. In kinetic analyses, Keap1-DLGex binding follows a fast-association and fast-dissociation model, while Keap1-ETGE binding contains a slow-reaction step that leads to a stable conformation. These results demonstrate that the mode of DLGex binding to Keap1 is distinct from that of ETGE structurally, thermodynamically, and kinetically and support our contention that the DLGex motif serves as a converter transmitting environmental stress to Nrf2 induction as the latch site.
- Department of Medical Biochemistry, Tohoku University Graduate School of Medicine, Sendai, Japan.
Organizational Affiliation: