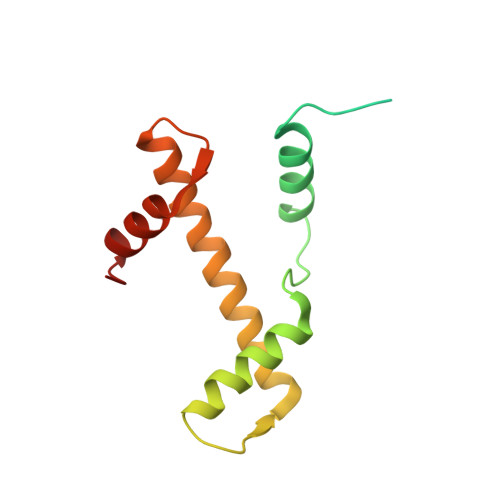
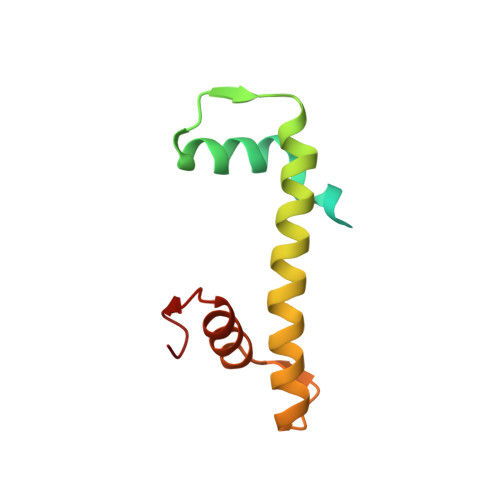
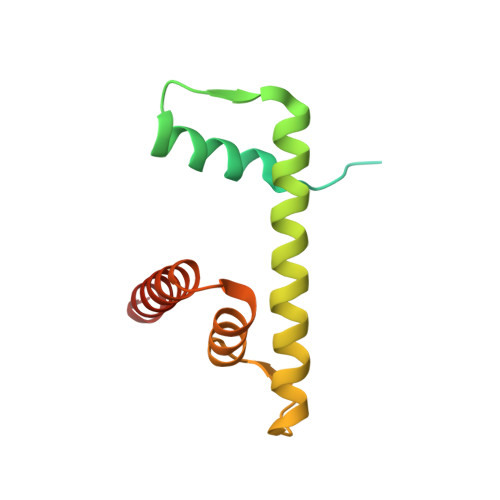
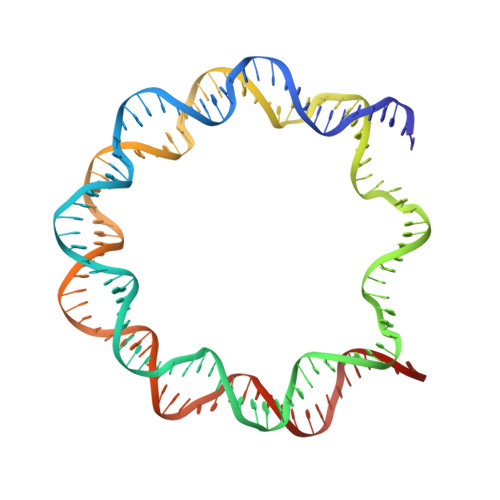
Structure of human nucleosome containing the testis-specific histone variant TSH2B.
Urahama, T., Horikoshi, N., Osakabe, A., Tachiwana, H., Kurumizaka, H.(2014) Acta Crystallogr F Struct Biol Commun 70: 444-449
- PubMed: 24699735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14004695
- Primary Citation Related Structures:
3WKJ - PubMed Abstract:
The human histone H2B variant TSH2B is highly expressed in testis and may function in the chromatin transition during spermatogenesis. In the present study, the crystal structure of the human testis-specific nucleosome containing TSH2B was determined at 2.8 Å resolution. A local structural difference between TSH2B and canonical H2B in nucleosomes was detected around the TSH2B-specific amino-acid residue Ser85. The TSH2B Ser85 residue does not interact with H4 in the nucleosome, but in the canonical nucleosome the H2B Asn84 residue (corresponding to the TSH2B Ser85 residue) forms water-mediated hydrogen bonds with the H4 Arg78 residue. In contrast, the other TSH2B-specific amino-acid residues did not induce any significant local structural changes in the TSH2B nucleosome. These findings may provide important information for understanding how testis-specific histone variants form nucleosomes during spermatogenesis.
- Laboratory of Structural Biology, Graduate School of Advanced Science and Engineering, Waseda University, 2-2 Wakamatsu-cho, Shinjuku-ku, Tokyo 162-8480, Japan.
Organizational Affiliation: