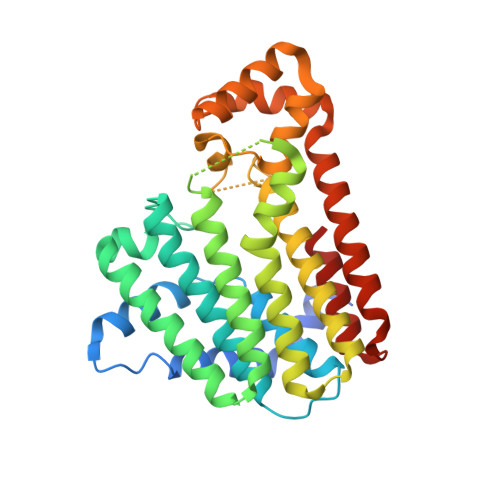
Crystal structures of ligand-bound octaprenyl pyrophosphate synthase from Escherichia coli reveal the catalytic and chain-length determining mechanisms.
Han, X., Chen, C.C., Kuo, C.J., Huang, C.H., Zheng, Y., Ko, T.P., Zhu, Z., Feng, X., Wang, K., Oldfield, E., Wang, A.H., Liang, P.H., Guo, R.T., Ma, Y.(2015) Proteins 83: 37-45
- PubMed: 24895191 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24618
- Primary Citation Related Structures:
3WJK, 3WJN, 3WJO - PubMed Abstract:
Octaprenyl pyrophosphate synthase (OPPs) catalyzes consecutive condensation reactions of one allylic substrate farnesyl pyrophosphate (FPP) and five homoallylic substrate isopentenyl pyrophosphate (IPP) molecules to form a C40 long-chain product OPP, which serves as a side chain of ubiquinone and menaquinone. OPPs belongs to the trans-prenyltransferase class of proteins. The structures of OPPs from Escherichia coli were solved in the apo-form as well as in complexes with IPP and a FPP thio-analog, FsPP, at resolutions of 2.2-2.6 Å, and revealed the detailed interactions between the ligands and enzyme. At the bottom of the active-site tunnel, M123 and M135 act in concert to form a wall which determines the final chain length. These results represent the first ligand-bound crystal structures of a long-chain trans-prenyltransferase and provide new information on the mechanisms of catalysis and product chain elongation.
- Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin, China.
Organizational Affiliation: