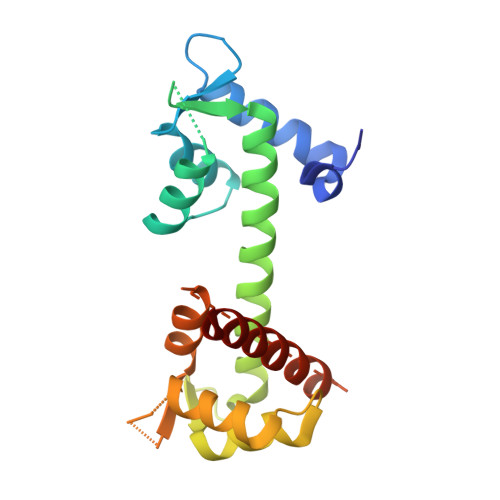
Structural Basis for the Modulation of the Neuronal Voltage-Gated Sodium Channel NaV1.6 by Calmodulin
Chichili, V.P.R., Xiao, Y., Seetharaman, J., Cummins, T.R., Sivaraman, J.(2013) Sci Rep 3: 2435-2435
- PubMed: 23942337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep02435
- Primary Citation Related Structures:
3WFN - PubMed Abstract:
The neuronal-voltage gated sodium channel (VGSC), Na(V)1.6, plays an important role in propagating action potentials along myelinated axons. Calmodulin (CaM) is known to modulate the inactivation kinetics of Na(V)1.6 by interacting with its IQ motif. Here we report the crystal structure of apo-CaM:Na(V)1.6IQ motif, along with functional studies. The IQ motif of Na(V)1.6 adopts an α-helical conformation in its interaction with the C-lobe of CaM. CaM uses different residues to interact with Na(V)1.6IQ motif depending on the presence or absence of Ca²⁺. Three residues from Na(V)1.6, Arg1902, Tyr1904 and Arg1905 were identified as the key common interacting residues in both the presence and absence of Ca²⁺. Substitution of Arg1902 and Tyr1904 with alanine showed a reduced rate of Na(V)1.6 inactivation in electrophysiological experiments in vivo. Compared with other CaM:Na(V) complexes, our results reveal a different mode of interaction for CaM:Na(V)1.6 and provides structural insight into the isoform-specific modulation of VGSCs.
- Department of Biological Sciences, National University of Singapore, Singapore, 117543.
Organizational Affiliation: