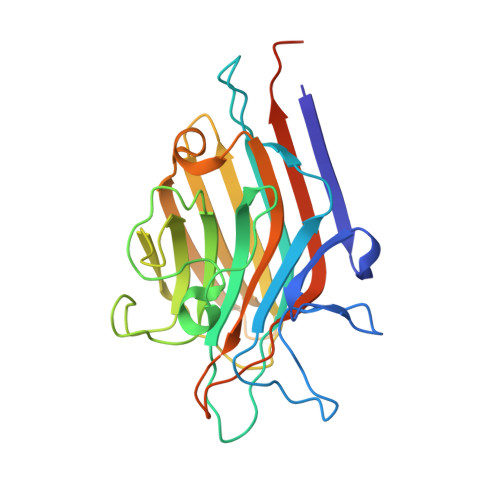
Phytohemagglutinin from Phaseolus vulgaris (PHA-E) displays a novel glycan recognition mode using a common legume lectin fold
Nagae, M., Soga, K., Morita-Matsumoto, K., Hanashima, S., Ikeda, A., Yamamoto, K., Yamaguchi, Y.(2014) Glycobiology 24: 368-378
- PubMed: 24436051 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwu004
- Primary Citation Related Structures:
3WCR, 3WCS, 3WOG - PubMed Abstract:
Phytohemagglutinin from Phaseolus vulgaris (PHA-E), a legume lectin, has an unusual specificity toward biantennary galactosylated N-glycan with bisecting N-acetylglucosamine (GlcNAc). To investigate the interaction in detail, we have solved the crystal structures of PHA-E without ligand and in complex with biantennary N-glycan derivatives. PHA-E interacts with the trisaccharide unit (Galβ1-4GlcNAcβ1-2Man) in a manner completely different from that of mannose/glucose-specific legume lectins. The inner mannose residue binds to a novel site on the protein, and its rotation is opposite to that occurring in the monosaccharide-binding site of other lectins around the sugar O3 axis. Saturation-transfer difference NMR using biantennary di-galactosylated and bisected glycans reveals that PHA-E interacts with both antennas almost equally. The unique carbohydrate interaction explains the glycan-binding specificity and high affinity.
- Structural Glycobiology Team, Systems Glycobiology Research Group, RIKEN-Max Planck Joint Research Center, RIKEN Global Research Cluster, 2-1 Hirosawa, Wako, Saitama 351-0198, Japan.
Organizational Affiliation: