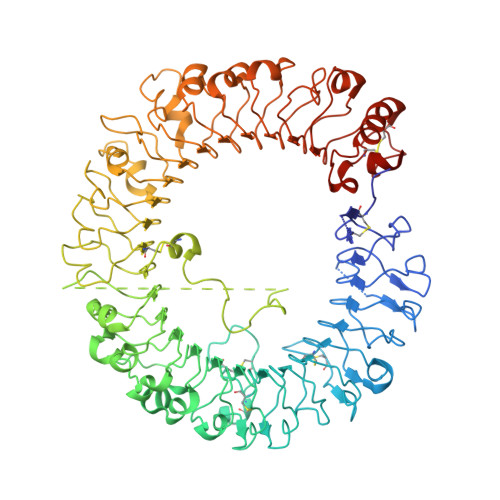
Structural reorganization of the Toll-like receptor 8 dimer induced by agonistic ligands
Tanji, H., Ohto, U., Shibata, T., Miyake, K., Shimizu, T.(2013) Science 339: 1426-1429
- PubMed: 23520111 Search on PubMed
- DOI: https://doi.org/10.1126/science.1229159
- Primary Citation Related Structures:
3W3G, 3W3J, 3W3K, 3W3L, 3W3M, 3W3N - PubMed Abstract:
Toll-like receptor 7 (TLR7) and TLR8 recognize single-stranded RNA and initiate innate immune responses. Several synthetic agonists of TLR7-TLR8 display novel therapeutic potential; however, the molecular basis for ligand recognition and activation of signaling by TLR7 or TLR8 is largely unknown. In this study, the crystal structures of unliganded and ligand-induced activated human TLR8 dimers were elucidated. Ligand recognition was mediated by a dimerization interface formed by two protomers. Upon ligand stimulation, the TLR8 dimer was reorganized such that the two C termini were brought into proximity. The loop between leucine-rich repeat 14 (LRR14) and LRR15 was cleaved; however, the N- and C-terminal halves remained associated and contributed to ligand recognition and dimerization. Thus, ligand binding induces reorganization of the TLR8 dimer, which enables downstream signaling processes.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: