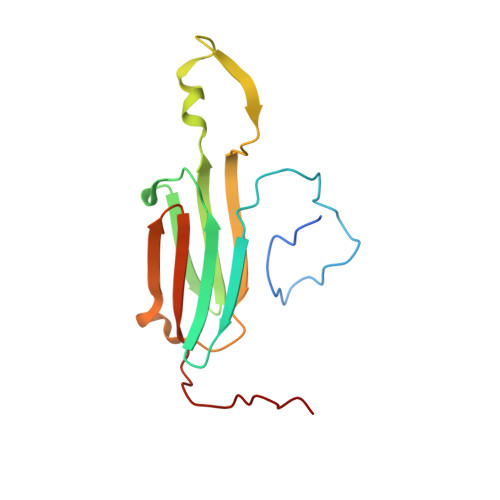
Nonequivalence Observed for the 16-Meric Structure of a Small Heat Shock Protein, SpHsp16.0, from Schizosaccharomyces pombe
Hanazono, Y., Takeda, K., Oka, T., Abe, T., Tomonari, T., Akiyama, N., Aikawa, Y., Yohda, M., Miki, K.(2013) Structure 21: 220-228
- PubMed: 23273429
- DOI: https://doi.org/10.1016/j.str.2012.11.015
- Primary Citation Related Structures:
3W1Z - PubMed Abstract:
Small heat shock proteins (sHsps) play a role in preventing the fatal aggregation of denatured proteins in the presence of stresses. The sHsps exist as monodisperse oligomers in their resting state. Because the hydrophobic N-terminal regions of sHsps are possible interaction sites for denatured proteins, the manner of assembly of the oligomer is critical for the activation and inactivation mechanisms. Here, we report the oligomer architecture of SpHsp16.0 from Schizosaccharomyces pombe determined with X-ray crystallography and small angle X-ray scattering. Both results indicate that eight dimers of SpHsp16.0 form an elongated sphere with 422 symmetry. The monomers show nonequivalence in the interaction with neighboring monomers and conformations of the N- and C-terminal regions. Variants for the N-terminal phenylalanine residues indicate that the oligomer formation ability is highly correlated with chaperone activity. Structural and biophysical results are discussed in terms of their possible relevance to the activation mechanism of SpHsp16.0.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: