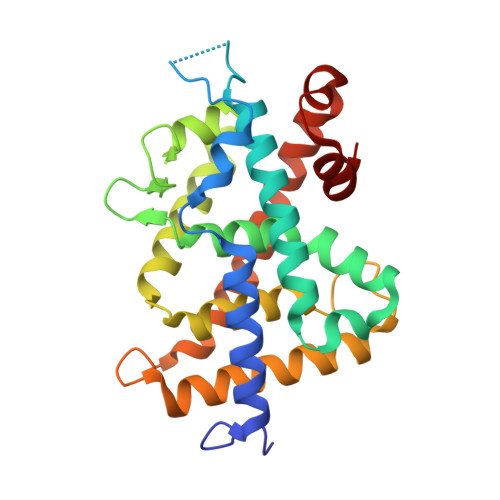
Structural basis for vitamin D receptor agonism by novel non-secosteroidal ligands.
Asano, L., Ito, I., Kuwabara, N., Waku, T., Yanagisawa, J., Miyachi, H., Shimizu, T.(2013) FEBS Lett 587: 957-963
- PubMed: 23462137 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2013.02.028
- Primary Citation Related Structures:
3W0G, 3W0H, 3W0I, 3W0J - PubMed Abstract:
Non-secosteroidal ligands for vitamin D receptor (VDR) have been developed for the agonist with non-calcemic profiles. Here, we provide the structural mechanism of VDR agonism by novel non-secosteroidal ligands. All ligands had the similar efficacy, while two had the higher potency. Crystallographic analyses revealed that all ligands interacted with helix H10 and the loop between helices H6 and H7 in a similar manner, but also that the two ligands with higher potency had different interaction modes. This study suggests that distinct ligand potency depend upon differences in the formation and rearrangement of hydrogen-bond networks induced by each ligand.
- Graduate School of Pharmaceutical Sciences, University of Tokyo, Hongo, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: