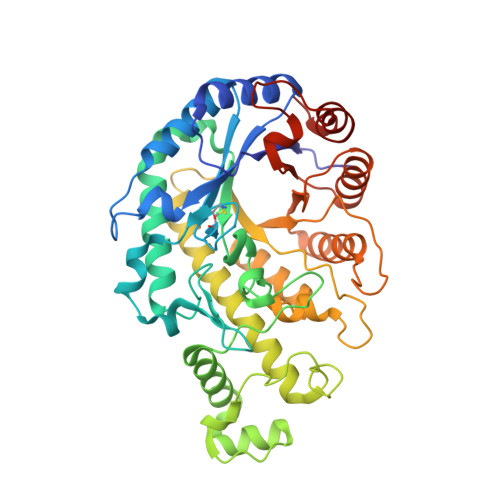
Structural analysis by X-ray crystallography and small-angle scattering of the multi-domain beta-amylase from Paenibacillus polymyxa
Nishimura, S., Fujioka, T., Takahashi, R., Nakaniwa, T., Fukada, H., Inui, T., Tada, T., Kawaguchi, T., Sumitani, J.To be published.