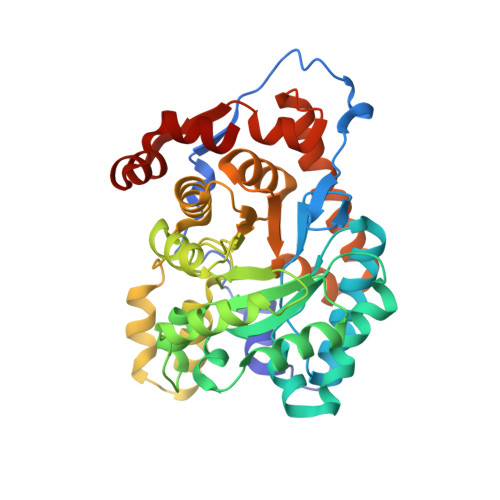
Substrate-Induced Change in the Quaternary Structure of Type 2 Isopentenyl Diphosphate Isomerase from Sulfolobus shibatae.
Nakatani, H., Goda, S., Unno, H., Nagai, T., Yoshimura, T., Hemmi, H.(2012) J Bacteriol 194: 3216-3224
- PubMed: 22505674 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00068-12
- Primary Citation Related Structures:
3VKJ - PubMed Abstract:
Type 2 isopentenyl diphosphate isomerase catalyzes the interconversion between two active units for isoprenoid biosynthesis, i.e., isopentenyl diphosphate and dimethylallyl diphosphate, in almost all archaea and in some bacteria, including human pathogens. The enzyme is a good target for discovery of antibiotics because it is essential for the organisms that use only the mevalonate pathway to produce the active isoprene units and because humans possess a nonhomologous isozyme, type 1 isopentenyl diphosphate isomerase. However, type 2 enzymes were reportedly inhibited by mechanism-based drugs for the type 1 enzyme due to their surprisingly similar reaction mechanisms. Thus, a different approach is now required to develop new inhibitors specific to the type 2 enzyme. X-ray crystallography and gel filtration chromatography revealed that the enzyme from a thermoacidophilic archaeon, Sulfolobus shibatae, is in the octameric state at a high concentration. Interestingly, a part of the regions that are involved in the substrate binding in the previously reported tetrameric structures is integral to the formation of the tetramer-tetramer interface in the substrate-free octameric structure. Site-directed mutagenesis at such regions resulted in stabilization of the tetramer. Small-angle X-ray scattering, tryptophan fluorescence, and dynamic light scattering analyses showed that substrate binding causes the dissociation of an octamer into tetramers. This property, i.e., incompatibility between octamer formation and substrate binding, might provide clues to develop new specific inhibitors of the archaeal enzyme.
- Department of Applied Molecular Bioscience, Graduate School of Bioagricultural Sciences, Nagoya University, Nagoya, Aichi, Japan.
Organizational Affiliation: