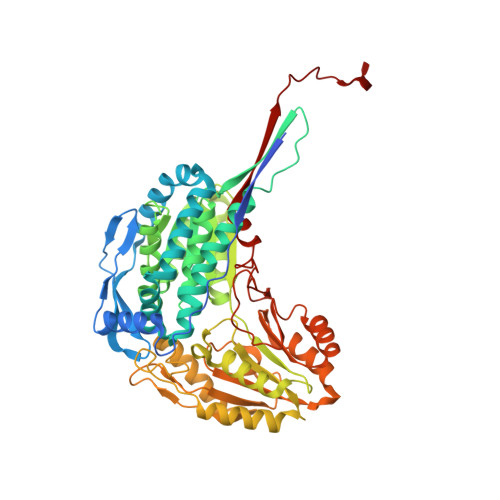
The Three-Dimensional Structural Basis of Type II Hyperprolinemia.
Srivastava, D., Singh, R.K., Moxley, M.A., Henzl, M.T., Becker, D.F., Tanner, J.J.(2012) J Mol Biology 420: 176-189
- PubMed: 22516612
- DOI: https://doi.org/10.1016/j.jmb.2012.04.010
- Primary Citation of Related Structures:
3V9G, 3V9H, 3V9I, 3V9J, 3V9K, 3V9L - PubMed Abstract:
Type II hyperprolinemia is an autosomal recessive disorder caused by a deficiency in Δ(1)-pyrroline-5-carboxylate dehydrogenase (P5CDH; also known as ALDH4A1), the aldehyde dehydrogenase that catalyzes the oxidation of glutamate semialdehyde to glutamate. Here, we report the first structure of human P5CDH (HsP5CDH) and investigate the impact of the hyperprolinemia-associated mutation of Ser352 to Leu on the structure and catalytic properties of the enzyme. The 2. 5-Å-resolution crystal structure of HsP5CDH was determined using experimental phasing. Structures of the mutant enzymes S352A (2.4 Å) and S352L (2.85 Å) were determined to elucidate the structural consequences of altering Ser352. Structures of the 93% identical mouse P5CDH complexed with sulfate ion (1.3 Å resolution), glutamate (1.5 Å), and NAD(+) (1.5 Å) were determined to obtain high-resolution views of the active site. Together, the structures show that Ser352 occupies a hydrophilic pocket and is connected via water-mediated hydrogen bonds to catalytic Cys348. Mutation of Ser352 to Leu is shown to abolish catalytic activity and eliminate NAD(+) binding. Analysis of the S352A mutant shows that these functional defects are caused by the introduction of the nonpolar Leu352 side chain rather than the removal of the Ser352 hydroxyl. The S352L structure shows that the mutation induces a dramatic 8-Å rearrangement of the catalytic loop. Because of this conformational change, Ser349 is not positioned to interact with the aldehyde substrate, conserved Glu447 is no longer poised to bind NAD(+), and Cys348 faces the wrong direction for nucleophilic attack. These structural alterations render the enzyme inactive.
- Department of Chemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: