Conformational Locking upon Cooperative Assembly of Notch Transcription Complexes.
Choi, S.H., Wales, T.E., Nam, Y., O'Donovan, D.J., Sliz, P., Engen, J.R., Blacklow, S.C.(2012) Structure 20: 340-349
- PubMed: 22325781 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.12.011
- Primary Citation Related Structures:
3V79 - PubMed Abstract:
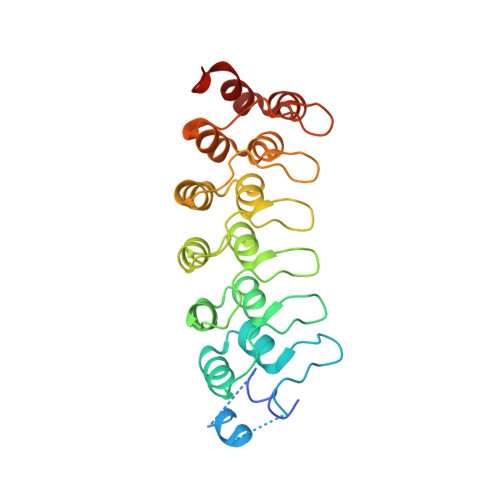
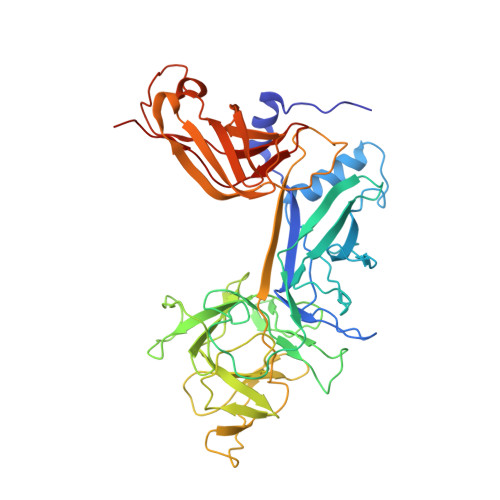
The Notch intracellular domain (NICD) forms a transcriptional activation complex with the DNA-binding factor CSL and a transcriptional co-activator of the Mastermind family (MAML). The "RAM" region of NICD recruits Notch to CSL, facilitating the binding of MAML at the interface between the ankyrin (ANK) repeat domain of NICD and CSL. Here, we report the X-ray structure of a human MAML1/RAM/ANK/CSL/DNA complex, and probe changes in component dynamics upon stepwise assembly of a MAML1/NICD/CSL complex using HX-MS. Association of CSL with NICD exerts remarkably little effect on the exchange kinetics of the ANK domain, whereas MAML1 binding greatly retards the exchange kinetics of ANK repeats 2-3. These exchange patterns identify critical features contributing to the cooperative assembly of Notch transcription complexes (NTCs), highlight the importance of MAML recruitment in rigidifying the ANK domain and stabilizing its interface with CSL, and rationalize the requirement for MAML1 in driving cooperative dimerization of NTCs on paired-site DNA.
- Department of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA 02115, USA.
Organizational Affiliation: