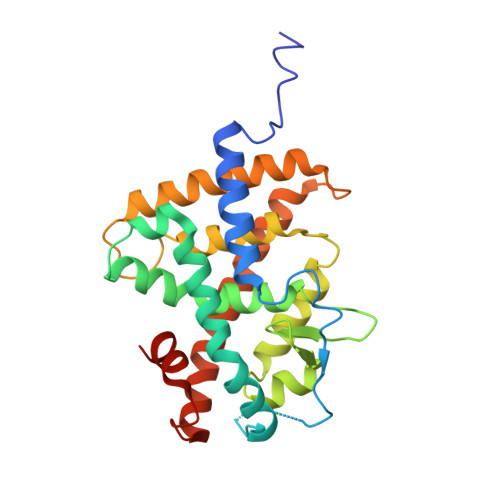
Structural basis for negative cooperativity within agonist-bound TR:RXR heterodimers.
Putcha, B.D., Wright, E., Brunzelle, J.S., Fernandez, E.J.(2012) Proc Natl Acad Sci U S A 109: 6084-6087
- PubMed: 22474364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1119852109
- Primary Citation Related Structures:
3UVV - PubMed Abstract:
Thyroid hormones such as 3,3',5 triiodo-L-thyronine (T3) control numerous aspects of mammalian development and metabolism. The actions of such hormones are mediated by specific thyroid hormone receptors (TRs). TR belongs to the nuclear receptor family of modular transcription factors that binds to specific DNA-response elements within target promoters. These receptors can function as homo- or heterodimers such as TR:9-cis retinoic acid receptor (RXR). Here, we present the atomic resolution structure of the TRα•T3:RXRα•9-cis retinoic acid (9c) ligand binding domain heterodimer complex at 2.95 Å along with T3 hormone binding and dissociation and coactivator binding studies. Our data provide a structural basis for allosteric communication between T3 and 9c and negative cooperativity between their binding pockets. In this structure, both TR and RXR are in the active state conformation for optimal binding to coactivator proteins. However, the structure of TR•T3 within TR•T3:RXR•9c is in a relative state of disorder, and the observed kinetics of binding show that T3 dissociates more rapidly from TR•T3:RXR•9c than from TR•T3:RXR. Also, coactivator binding studies with a steroid receptor coactivator-1 (receptor interaction domains 1-3) fragment show lower affinities (K(a)) for TR•T3:RXR•9c than TR•T3:RXR. Our study corroborates previously reported observations from cell-based and binding studies and offers a structural mechanism for the repression of TR•T3:RXR transactivation by RXR agonists. Furthermore, the recent discoveries of multiple endogenous RXR agonists that mediate physiological tasks such as lipid biosynthesis underscore the pharmacological importance of negative cooperativity in ligand binding within TR:RXR heterodimers.
- University of Tennessee, Knoxville, TN 37996, USA.
Organizational Affiliation: