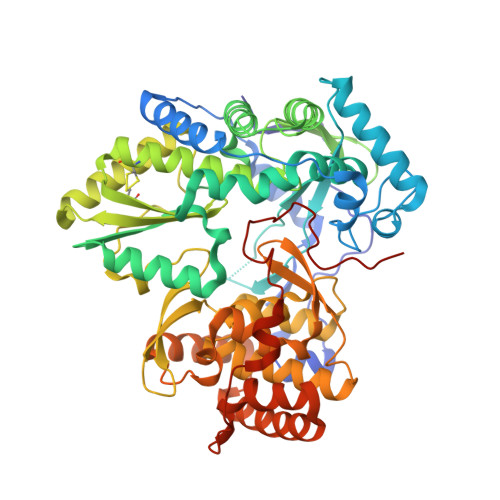
Synthesis of New 4,5-Dihydrofuranoindoles and Their Evaluation as HCV NS5B Polymerase Inhibitors.
Velazquez, F., Venkatraman, S., Lesburg, C.A., Duca, J., Rosenblum, S.B., Kozlowski, J.A., Njoroge, F.G.(2012) Org Lett 14: 556-559
- PubMed: 22220815 Search on PubMed
- DOI: https://doi.org/10.1021/ol203177g
- Primary Citation Related Structures:
3UPH, 3UPI - PubMed Abstract:
The synthesis of substituted 3,4-dihydrofuranoindoles is reported. These new indole compounds were used to synthesize potent HCV NS5B inhibitors. The binding mode of the dihydrofuranoindole-derived inhibitors was established via X-ray crystallographic studies.
- Merck Research Laboratories, 2015 Galloping Hill Road, Kenilworth, New Jersey 07033-1300, USA. francisco.velazquez@merck.com
Organizational Affiliation: