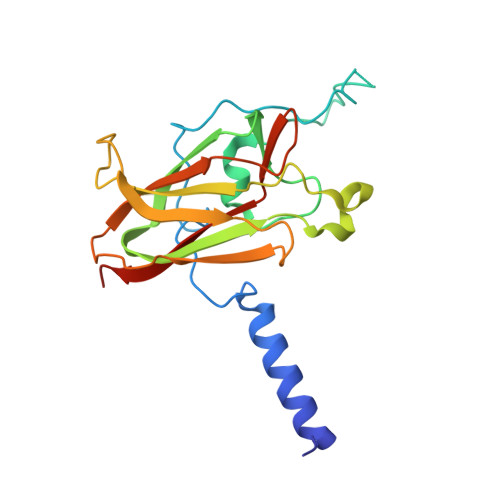
Structure of Sad1-UNC84 homology (SUN) domain defines features of molecular bridge in nuclear envelope
Zhou, Z.C., Du, X., Cai, Z., Song, X., Zhang, H., Mizuno, T., Suzuki, E., Yee, M.R., Berezov, A., Murali, R., Wu, S.-L., Karger, B.L., Greene, M.I., Wang, Q.(2012) J Biological Chem 287: 5317-5326
- PubMed: 22170055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.304543
- Primary Citation Related Structures:
3UNP - PubMed Abstract:
The SUN (Sad1-UNC-84 homology) domain is conserved in a number of nuclear envelope proteins involved in nuclear migration, meiotic telomere tethering, and antiviral responses. The LINC (linker of nucleoskeleton and cytoskeleton) complex, formed by the SUN and the nesprin proteins at the nuclear envelope, serves as a mechanical linkage across the nuclear envelope. Here we report the crystal structure of the SUN2 protein SUN domain, which reveals a homotrimer. The SUN domain is sufficient to mediate binding to the KASH (Klarsicht, ANC-1, and Syne homology) domain of nesprin 2, and the regions involved in the interaction have been identified. Binding of the SUN domain to the KASH domain is abolished by deletion of a region important for trimerization or by point mutations associated with nuclear migration failure. We propose a model of the LINC complex, where the SUN and the KASH domains form a higher ordered oligomeric network in the nuclear envelope. These findings provide the structural basis for understanding the function and the regulation of the LINC complex.
- Department of Pathology and Laboratory Medicine, University of Pennsylvania, Perelman School of Medicine, Philadelphia, Pennsylvania 19104, USA.
Organizational Affiliation: