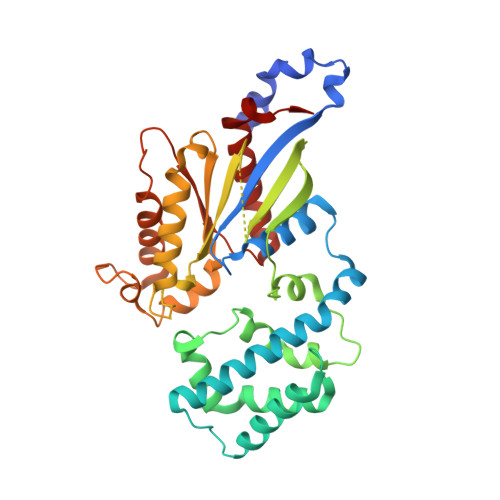
Structural Determinants Underlying the Temperature-sensitive Nature of a G-alpha Mutant in Asymmetric Cell Division of Caenorhabditis elegans
Johnston, C.A., Afshar, K., Snyder, J.T., Tall, G.G., Gonczy, P., Siderovski, D.P., Willard, F.S.To be published.