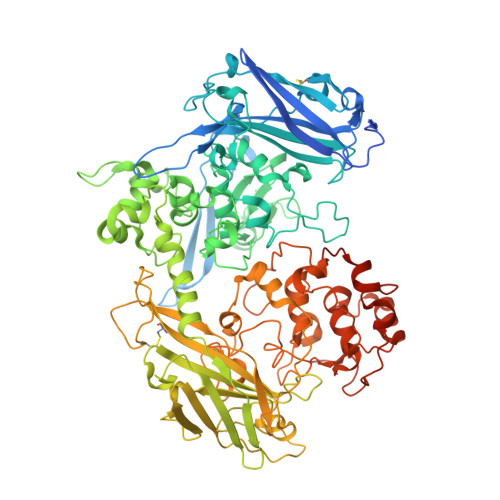
The Crystal Structure of the Extracellular 11-heme Cytochrome UndA Reveals a Conserved 10-heme Motif and Defined Binding Site for Soluble Iron Chelates.
Edwards, M.J., Hall, A., Shi, L., Fredrickson, J.K., Zachara, J.M., Butt, J.N., Richardson, D.J., Clarke, T.A.(2012) Structure 20: 1275-1284
- PubMed: 22682743 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.04.016
- Primary Citation Related Structures:
3UCP, 3UFH, 3UFK - PubMed Abstract:
Members of the genus Shewanella translocate deca- or undeca-heme cytochromes to the external cell surface thus enabling respiration using extracellular minerals and polynuclear Fe(III) chelates. The high resolution structure of the first undeca-heme outer membrane cytochrome, UndA, reveals a crossed heme chain with four potential electron ingress/egress sites arranged within four domains. Sequence and structural alignment of UndA and the deca-heme MtrF reveals the extra heme of UndA is inserted between MtrF hemes 6 and 7. The remaining UndA hemes can be superposed over the heme chain of the decaheme MtrF, suggesting that a ten heme core is conserved between outer membrane cytochromes. The UndA structure has also been crystallographically resolved in complex with substrates, an Fe(III)-nitrilotriacetate dimer or an Fe(III)-citrate trimer. The structural resolution of these UndA-Fe(III)-chelate complexes provides a rationale for previous kinetic measurements on UndA and other outer membrane cytochromes.
- Centre for Molecular and Structural Biochemistry, School of Biological Sciences and School of Chemistry, University of East Anglia, Norwich Research Park, Norwich NR4 7TJ, UK.
Organizational Affiliation: