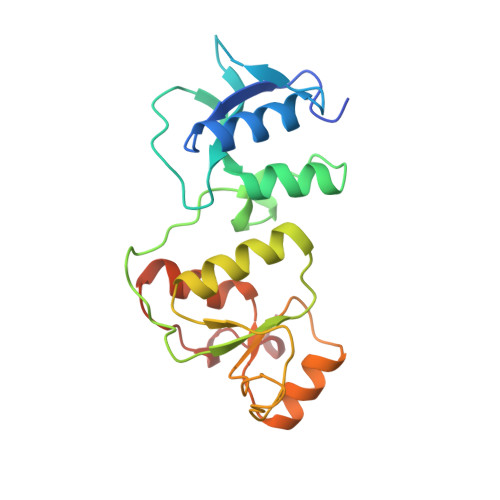
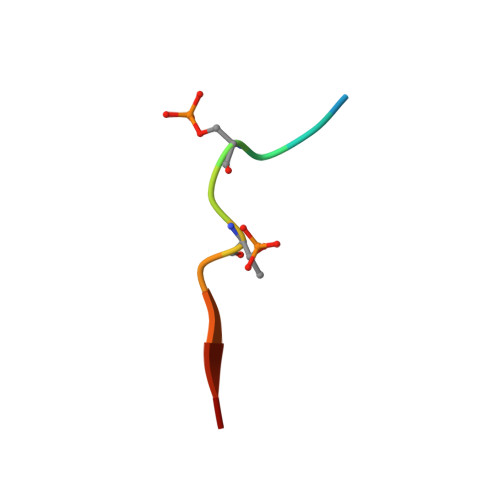
Structural insights into recognition of MDC1 by TopBP1 in DNA replication checkpoint control.
Leung, C.C., Sun, L., Gong, Z., Burkat, M., Edwards, R., Assmus, M., Chen, J., Glover, J.N.(2013) Structure 21: 1450-1459
- PubMed: 23891287 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.06.015
- Primary Citation Related Structures:
3UEN, 3UEO - PubMed Abstract:
Activation of the DNA replication checkpoint by the ATR kinase requires protein interactions mediated by the ATR-activating protein, TopBP1. Accumulation of TopBP1 at stalled replication forks requires the interaction of TopBP1 BRCT5 with the phosphorylated SDT repeats of the adaptor protein MDC1. Here, we present the X-ray crystal structures of the tandem BRCT4/5 domains of TopBP1 free and in complex with a MDC1 consensus pSDpT phosphopeptide. TopBP1 BRCT4/5 adopts a variant BRCT-BRCT packing interface and recognizes its target peptide in a manner distinct from that observed in previous tandem BRCT- peptide structures. The phosphate-binding pocket and positively charged residues in a variant loop in BRCT5 present an extended binding surface for the negatively charged MDC1 phosphopeptide. Mutations in this surface reduce binding affinity and recruitment of TopBP1 to γH2AX foci in cells. These studies reveal a different mode of phosphopeptide binding by BRCT domains in the DNA damage response.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta T6G 2H7, Canada.
Organizational Affiliation: