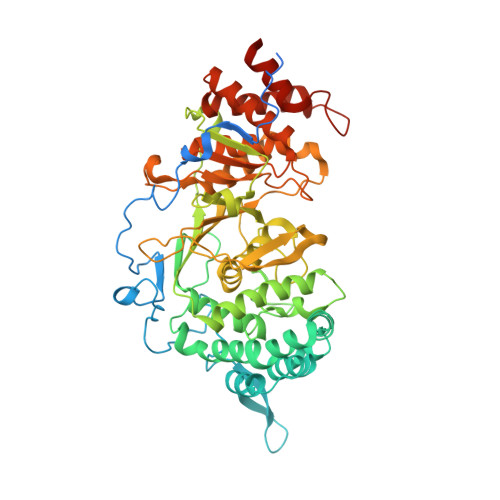
Structure of mammalian poly(ADP-ribose) glycohydrolase reveals a flexible tyrosine clasp as a substrate-binding element.
Kim, I.K., Kiefer, J.R., Ho, C.M., Stegeman, R.A., Classen, S., Tainer, J.A., Ellenberger, T.(2012) Nat Struct Mol Biol 19: 653-656
- PubMed: 22609859 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2305
- Primary Citation Related Structures:
3UEK, 3UEL - PubMed Abstract:
Reversible post-translational modification by poly(ADP-ribose) (PAR) regulates chromatin structure, DNA repair and cell fate in response to genotoxic stress. PAR glycohydrolase (PARG) removes PAR chains from poly ADP-ribosylated proteins to restore protein function and release oligo(ADP-ribose) chains to signal damage. Here we report crystal structures of mammalian PARG and its complex with a substrate mimic that reveal an open substrate-binding site and a unique 'tyrosine clasp' enabling endoglycosidic cleavage of branched PAR chains.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri, USA.
Organizational Affiliation: