Structure of the complex between CbpA J-domain and CbpM provides a link between chaperone and transcription regulation in bacterial heat shock response
Sarraf, N.S., Shi, R., Zhang, L., Cygler, M., Ekiel, I.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Chaperone-modulator protein CbpM | 102 | Klebsiella variicola 342 | Mutation(s): 0 Gene Names: cbpM, KPK_5158 | 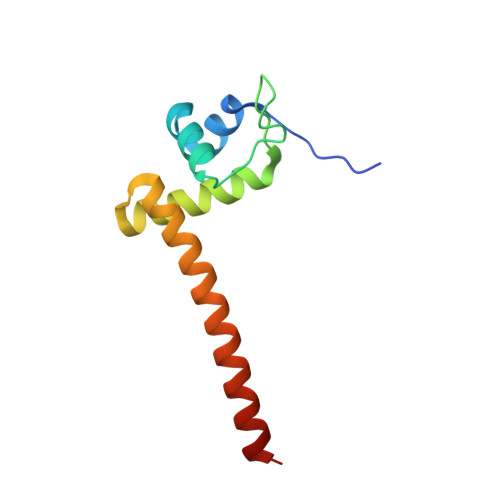 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | B5Y388 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Curved DNA-binding protein | 74 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: b1000, cbpA, JW0985 | 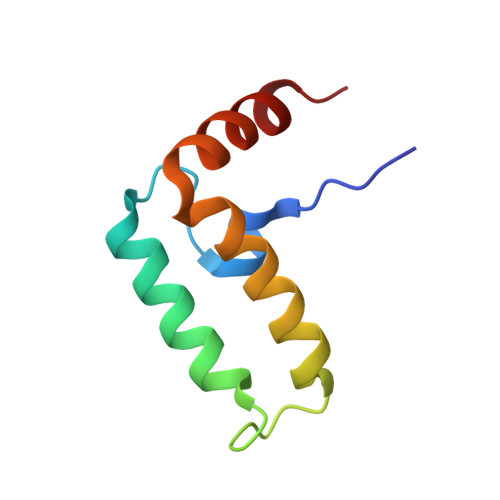 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P36659 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 52.816 | α = 90 |
| b = 77.094 | β = 90 |
| c = 111.231 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| SHARP | phasing |