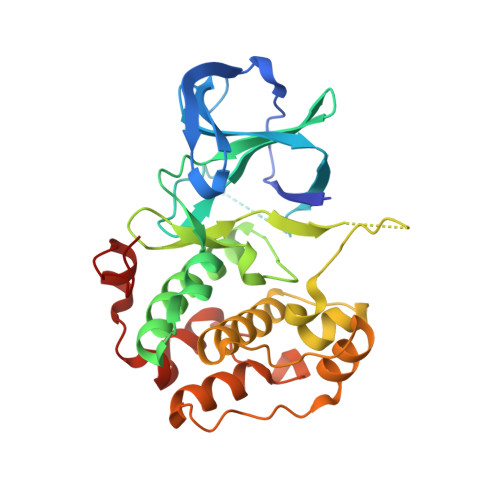
Insights into the Inhibition of the p90 Ribosomal S6 Kinase (RSK) by the Flavonol Glycoside SL0101 from the 1.5 A Crystal Structure of the N-Terminal Domain of RSK2 with Bound Inhibitor.
Utepbergenov, D., Derewenda, U., Olekhnovich, N., Szukalska, G., Banerjee, B., Hilinski, M.K., Lannigan, D.A., Stukenberg, P.T., Derewenda, Z.S.(2012) Biochemistry 51: 6499-6510
- PubMed: 22846040 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi300620c
- Primary Citation Related Structures:
3UBD, 4EL9 - PubMed Abstract:
The p90 ribosomal S6 family of kinases (RSK) are potential drug targets, due to their involvement in cancer and other pathologies. There are currently only two known selective inhibitors of RSK, but the basis for selectivity is not known. One of these inhibitors is a naturally occurring kaempferol-α-L-diacetylrhamnoside, SL0101. Here, we report the crystal structure of the complex of the N-terminal kinase domain of the RSK2 isoform with SL0101 at 1.5 Å resolution. The refined atomic model reveals unprecedented structural reorganization of the protein moiety, as compared to the nucleotide-bound form. The entire N-lobe, the hinge region, and the αD-helix undergo dramatic conformational changes resulting in a rearrangement of the nucleotide binding site with concomitant formation of a highly hydrophobic pocket spatially suited to accommodate SL0101. These unexpected results will be invaluable in further optimization of the SL0101 scaffold as a promising lead for a novel class of kinase inhibitors.
- Department of Molecular Physiology and Biological Physics, University of Virginia, School of Medicine, Charlottesville, VA 22908, USA.
Organizational Affiliation: