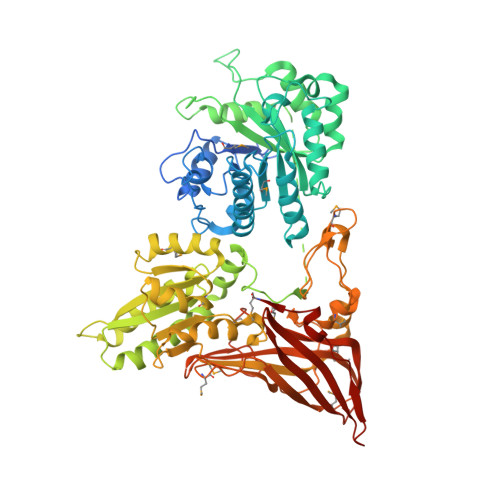
Structural insights into protein arginine symmetric dimethylation by PRMT5
Sun, L., Wang, M., Lv, Z., Yang, N., Liu, Y., Bao, S., Gong, W., Xu, R.M.(2011) Proc Natl Acad Sci U S A 108: 20538-20543
- PubMed: 22143770 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1106946108
- Primary Citation Related Structures:
3UA3, 3UA4 - PubMed Abstract:
Symmetric and asymmetric dimethylation of arginine are isomeric protein posttranslational modifications with distinct biological effects, evidenced by the methylation of arginine 3 of histone H4 (H4R3): symmetric dimethylation of H4R3 leads to repression of gene expression, while asymmetric dimethylation of H4R3 is associated with gene activation. The enzymes catalyzing these modifications share identifiable sequence similarities, but the relationship between their catalytic mechanisms is unknown. Here we analyzed the structure of a prototypic symmetric arginine dimethylase, PRMT5, and discovered that a conserved phenylalanine in the active site is critical for specifying symmetric addition of methyl groups. Changing it to a methionine significantly elevates the overall methylase activity, but also converts PRMT5 to an enzyme that catalyzes both symmetric and asymmetric dimethylation of arginine. Our results demonstrate a common catalytic mechanism intrinsic to both symmetric and asymmetric arginine dimethylases, and show that steric constrains in the active sites play an essential role in determining the product specificity of arginine methylases. This discovery also implies a potentially regulatable outcome of arginine dimethylation that may provide versatile control of eukaryotic gene expression.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: