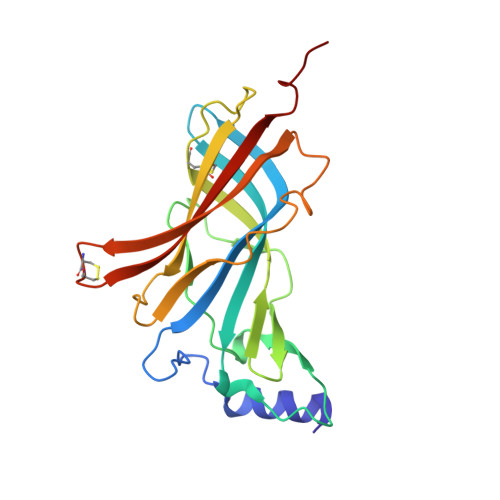
Intersubunit bridge formation governs agonist efficacy at nicotinic acetylcholine alpha 4 beta 2 receptors: unique role of halogen bonding revealed.
Rohde, L.A., Ahring, P.K., Jensen, M.L., Nielsen, E., Peters, D., Helgstrand, C., Krintel, C., Harpse, K., Gajhede, M., Kastrup, J.S., Balle, T.(2012) J Biological Chem 287: 4248-4259
- PubMed: 22170047 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.292243
- Primary Citation Related Structures:
3U8J, 3U8K, 3U8L, 3U8M, 3U8N - PubMed Abstract:
The α4β2 subtype of the nicotinic acetylcholine receptor has been pursued as a drug target for treatment of psychiatric and neurodegenerative disorders and smoking cessation aids for decades. Still, a thorough understanding of structure-function relationships of α4β2 agonists is lacking. Using binding experiments, electrophysiology and x-ray crystallography we have investigated a consecutive series of five prototypical pyridine-containing agonists derived from 1-(pyridin-3-yl)-1,4-diazepane. A correlation between binding affinities at α4β2 and the acetylcholine-binding protein from Lymnaea stagnalis (Ls-AChBP) confirms Ls-AChBP as structural surrogate for α4β2 receptors. Crystal structures of five agonists with efficacies at α4β2 from 21-76% were determined in complex with Ls-AChBP. No variation in closure of loop C is observed despite large efficacy variations. Instead, the efficacy of a compound appears tightly coupled to its ability to form a strong intersubunit bridge linking the primary and complementary binding interfaces. For the tested agonists, a specific halogen bond was observed to play a large role in establishing such strong intersubunit anchoring.
- Department of Medicinal Chemistry, Faculty of Pharmaceutical Sciences, University of Copenhagen, Copenhagen 2100, Denmark.
Organizational Affiliation: