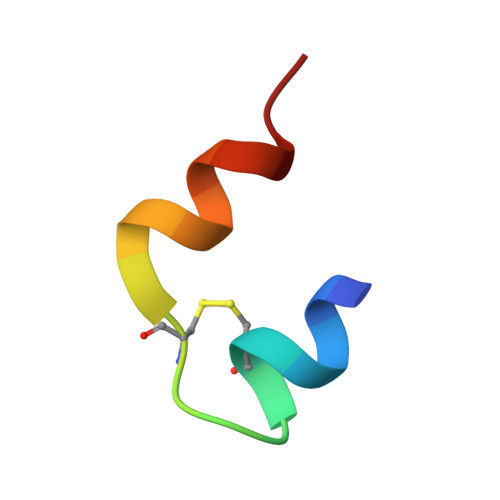
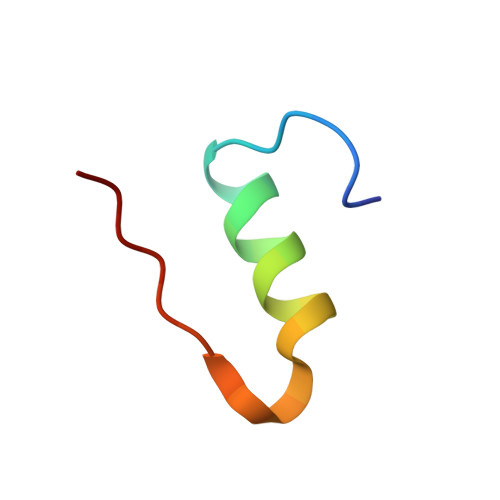
Novel covalently linked insulin dimer engineered to investigate the function of insulin dimerization.
Vinther, T.N., Norrman, M., Strauss, H.M., Huus, K., Schlein, M., Pedersen, T.A., Kjeldsen, T., Jensen, K.J., Hubalek, F.(2012) PLoS One 7: e30882-e30882
- PubMed: 22363506
- DOI: https://doi.org/10.1371/journal.pone.0030882
- Primary Citation of Related Structures:
3U4N - PubMed Abstract:
An ingenious system evolved to facilitate insulin binding to the insulin receptor as a monomer and at the same time ensure sufficient stability of insulin during storage. Insulin dimer is the cornerstone of this system. Insulin dimer is relatively weak, which ensures dissociation into monomers in the circulation, and it is stabilized by hexamer formation in the presence of zinc ions during storage in the pancreatic β-cell. Due to the transient nature of insulin dimer, direct investigation of this important form is inherently difficult. To address the relationship between insulin oligomerization and insulin stability and function, we engineered a covalently linked insulin dimer in which two monomers were linked by a disulfide bond. The structure of this covalent dimer was identical to the self-association dimer of human insulin. Importantly, this covalent dimer was capable of further oligomerization to form the structural equivalent of the classical hexamer. The covalently linked dimer neither bound to the insulin receptor, nor induced a metabolic response in vitro. However, it was extremely thermodynamically stable and did not form amyloid fibrils when subjected to mechanical stress, underlining the importance of oligomerization for insulin stability.
- Diabetes Research Unit, Novo Nordisk A/S, Novo Nordisk Park, Måløv, Denmark.
Organizational Affiliation: