Engineering and structural characterization of a linear polyubiquitin-specific antibody.
Matsumoto, M.L., Dong, K.C., Yu, C., Phu, L., Gao, X., Hannoush, R.N., Hymowitz, S.G., Kirkpatrick, D.S., Dixit, V.M., Kelley, R.F.(2012) J Mol Biology 418: 134-144
- PubMed: 22227388 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.12.053
- Primary Citation Related Structures:
3U30 - PubMed Abstract:
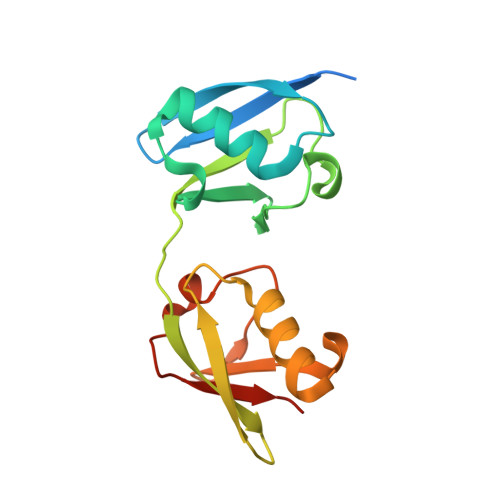
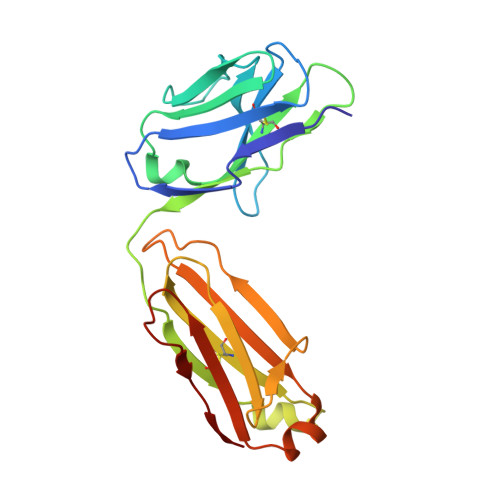
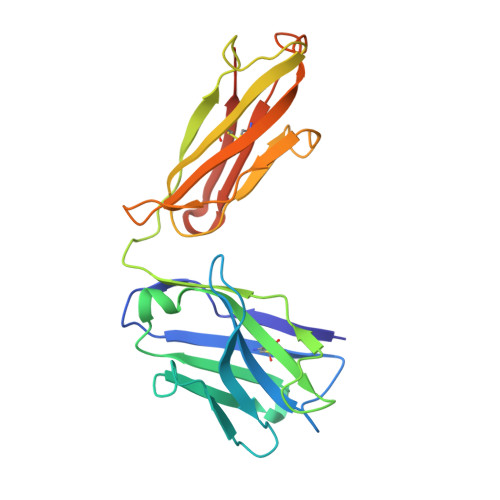
Polyubiquitination is an essential posttranslational modification that plays critical roles in cellular signaling. PolyUb (polyubiquitin) chains are formed by linking the carboxyl-terminus of one Ub (ubiquitin) subunit to either a lysine residue or the amino-terminus of an adjacent Ub. Linkage through the amino-terminus results in linear polyubiquitination that has recently been demonstrated to be a key step in nuclear factor κB activation; however, tools to study linear chains have been lacking. We therefore engineered a linear-linkage-specific antibody that is functional in Western blot, immunoprecipitation, and immunofluorescence applications. A crystal structure of the linear-linkage-specific antibody Fab fragment in complex with linear diubiquitin provides molecular insight into the nature of linear chain specificity. We use the antibody to demonstrate that linear polyUb is up-regulated upon tumor necrosis factor α stimulation of cells, consistent with a critical role in nuclear factor κB signaling. This antibody provides an essential tool for further investigation of the function of linear chains.
- Department of Antibody Engineering, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: