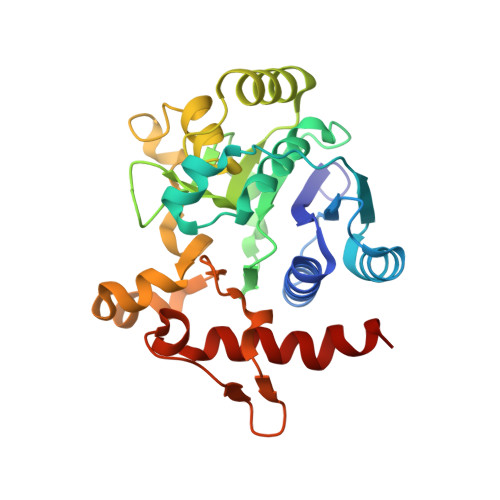
Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and 1'-deoxyglucose
Chaikuad, A., Froese, D.S., Krysztofinska, E., von Delft, F., Weigelt, J., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Oppermann, U., Yue, W.W., Structural Genomics Consortium (SGC)To be published.