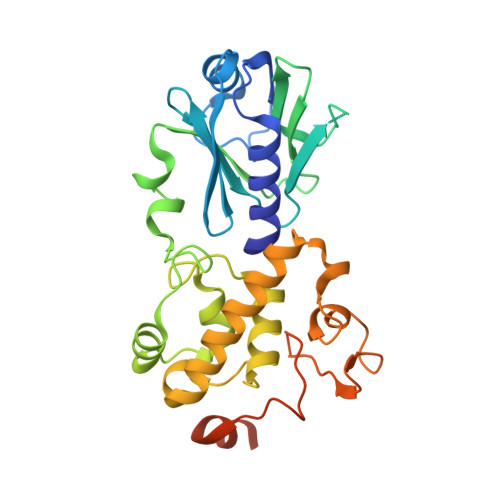
Structural and biochemical studies of a plant formamidopyrimidine-DNA glycosylase reveal why eukaryotic Fpg glycosylases do not excise 8-oxoguanine.
Duclos, S., Aller, P., Jaruga, P., Dizdaroglu, M., Wallace, S.S., Doublie, S.(2012) DNA Repair (Amst) 11: 714-725
- PubMed: 22789755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.dnarep.2012.06.004
- Primary Citation Related Structures:
3TWK, 3TWL, 3TWM - PubMed Abstract:
Formamidopyrimidine-DNA glycosylase (Fpg; MutM) is a DNA repair enzyme widely distributed in bacteria. Fpg recognizes and excises oxidatively modified purines, 4,6-diamino-5-formamidopyrimidine, 2,6-diamino-4-hydroxy-5-formamidopyrimidine and 8-oxoguanine (8-oxoG), with similar excision kinetics. It exhibits some lesser activity toward 8-oxoadenine. Fpg enzymes are also present in some plant and fungal species. The eukaryotic Fpg homologs exhibit little or no activity on DNA containing 8-oxoG, but they recognize and process its oxidation products, guanidinohydantoin (Gh) and spiroiminohydantoin (Sp). To date, several structures of bacterial Fpg enzymes unliganded or in complex with DNA containing a damaged base have been published but there is no structure of a eukaryotic Fpg. Here we describe the first crystal structure of a plant Fpg, Arabidopsis thaliana (AthFpg), unliganded and bound to DNA containing an abasic site analog, tetrahydrofuran (THF). Although AthFpg shares a common architecture with other Fpg glycosylases, it harbors a zincless finger, previously described in a subset of Nei enzymes, such as human NEIL1 and Mimivirus Nei1. Importantly the "αF-β9/10 loop" capping 8-oxoG in the active site of bacterial Fpg is very short in AthFpg. Deletion of a segment encompassing residues 213-229 in Escherichia coli Fpg (EcoFpg) and corresponding to the "αF-β9/10 loop" does not affect the recognition and removal of oxidatively damaged DNA base lesions, with the exception of 8-oxoG. Although the exact role of the loop remains to be further explored, it is now clear that this protein segment is specific to the processing of 8-oxoG.
- Department of Microbiology and Molecular Genetics, The Markey Center for Molecular Genetics, University of Vermont, Stafford Hall, Burlington, VT 05405-0068, USA. swallace@uvm.edu
Organizational Affiliation: