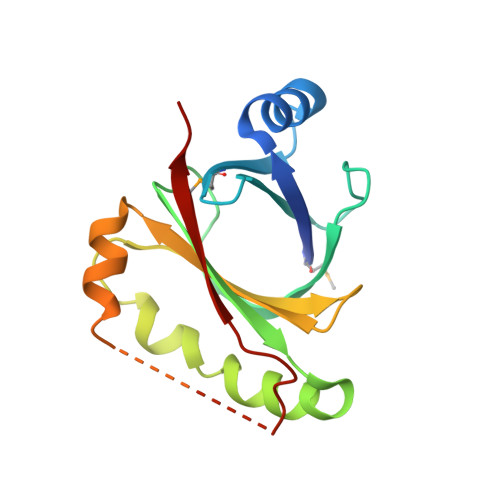
Bacillus subtilis HmoB is a heme oxygenase with a novel structure.
Park, S., Choi, S., Choe, J.(2012) BMB Rep 45: 239-241
- PubMed: 22531134 Search on PubMed
- DOI: https://doi.org/10.5483/bmbrep.2012.45.4.239
- Primary Citation Related Structures:
3TVZ - PubMed Abstract:
Iron availability is limited in the environment and most bacteria have developed a system to acquire iron from host hemoproteins. Heme oxygenase plays an important role by degrading heme group and releasing the essential nutrient iron. The structure of Bacillus subtilis HmoB was determined to 2.0 A resolution. B. subtilis HmoB contains a typical antibiotic biosynthesis monooxygenase (ABM) domain that spans from 71 to 146 residues and belongs to the IsdG family heme oxygenases. Comparison of HmoB and IsdG family proteins showed that the C-terminal region of HmoB has similar sequence and structure to IsdG family proteins and contains conserved critical residues for heme degradation. However, HmoB is distinct from other IsdG family proteins in that HmoB is about 60 amino acids longer in the N-terminus and does not form a dimer whereas previously studied IsdG family heme oxygenases form functional homodimers. Interestingly, the structure of monomeric HmoB resembles the dimeric structure of IsdG family proteins. Hence, B. subtilis HmoB is a heme oxygenase with a novel structural feature.
- Department of Life Science, University of Seoul, Seoul, Korea.
Organizational Affiliation: