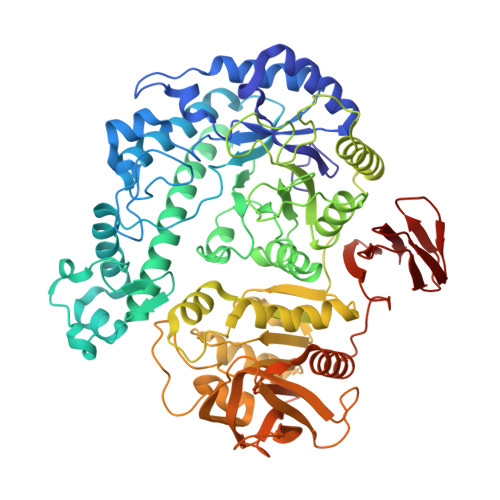
Structural analysis, enzymatic characterization, and catalytic mechanisms of beta-galactosidase from Bacillus circulans sp. alkalophilus.
Maksimainen, M., Paavilainen, S., Hakulinen, N., Rouvinen, J.(2012) FEBS J 279: 1788-1798
- PubMed: 22385475 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08555.x
- Primary Citation Related Structures:
3TTS, 3TTY - PubMed Abstract:
Crystal structures of native and α-D-galactose-bound Bacillus circulans sp. alkalophilus β-galactosidase (Bca-β-gal) were determined at 2.40 and 2.25 Å resolutions, respectively. Bca-β-gal is a member of family 42 of glycoside hydrolases, and forms a 460 kDa hexameric structure in crystal. The protein consists of three domains, of which the catalytic domain has an (α/β)(8) barrel structure with a cluster of sulfur-rich residues inside the β-barrel. The shape of the active site is clearly more open compared to the only homologous structure available in the Protein Data Bank. This is due to the number of large differences in the loops that connect the C-terminal ends of the β-strands to the N-terminal ends of the α-helices within the (α/β)(8) barrel. The complex structure shows that galactose binds to the active site as an α-anomer and induces clear conformational changes in the active site. The implications of α-D-galactose binding with respect to the catalytic mechanism are discussed. In addition, we suggest that β-galactosidases mainly utilize a reverse hydrolysis mechanism for synthesis of galacto-oligosaccharides.
- Department of Chemistry, University of Eastern Finland, Joensuu, Finland.
Organizational Affiliation: