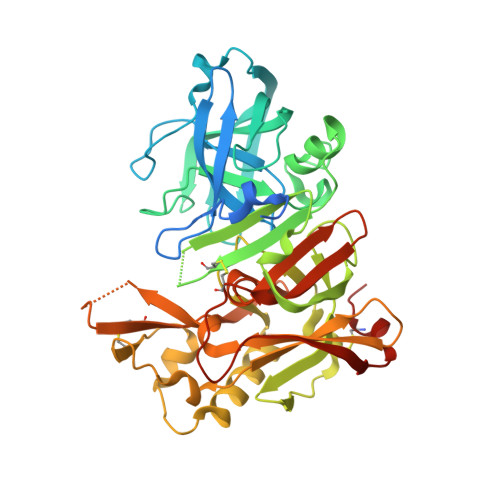
Flexibility of the flap in the active site of BACE1 as revealed by crystal structures and molecular dynamics simulations
Xu, Y.C., Li, M.J., Greenblatt, H., Chen, W.Y., Paz, A., Dym, O., Peleg, Y., Chen, T.T., Shen, X., He, J.H., Jiang, H.L., Silman, I., Sussman, J.L.(2012) Acta Crystallogr D Biol Crystallogr 68: 13-25
- PubMed: 22194329 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444911047251
- Primary Citation Related Structures:
3TPJ, 3TPL, 3TPP, 3TPR - PubMed Abstract:
β-Secretase (β-site amyloid precursor protein-cleaving enzyme 1; BACE1) is a transmembrane aspartic protease that cleaves the β-amyloid precursor protein en route to generation of the amyloid β-peptide (Aβ) that is believed to be responsible for the Alzheimer's disease amyloid cascade. It is thus a prime target for the development of inhibitors which may serve as drugs in the treatment and/or prevention of Alzheimer's disease. In the following determination of the crystal structures of both apo and complexed BACE1, structural analysis of all crystal structures of BACE1 deposited in the PDB and molecular dynamics (MD) simulations of monomeric and `dimeric' BACE1 were used to study conformational changes in the active-site region of the enzyme. It was observed that a flap able to cover the active site is the most flexible region, adopting multiple conformational states in the various crystal structures. Both the presence or absence of an inhibitor within the active site and the crystal packing are shown to influence the flap's conformation. An open conformation of the flap is mostly observed in the apo structures, while direct hydrogen-bonding interaction between main-chain atoms of the flap and the inhibitor is a prerequisite for the flap to adopt a closed conformation in the crystal structures of complexes. Thus, a systematic study of the conformational flexibility of the enzyme may not only contribute to structure-based drug design of BACE1 inhibitors and of other targets with flexible conformations, but may also help to better understand the mechanistic events associated with the binding of substrates and inhibitors to the enzyme.
- Drug Discovery and Design Center, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, People's Republic of China. ycxu@mail.shcnc.ac.cn
Organizational Affiliation: